Main Page
Welcome to the CGD Public Wiki
Seminal Candida Papers
Strain Information
Candida albicans Strains
SC5314
Genotype: wild type
Notes: Wild-type strain used in the systematic sequencing project, the reference sequence stored in CGD. The original strain background from which most of the common laboratory strains are derived. This strain is virulent in a mouse model of systemic infection and is frequently used as a wild-type control. In their 2004 Genome Biology paper on C. albicans genome sequence, Frank Odds, Al Brown and Neil Gow explain the origins of SC5314: "Strain SC5314 was used in the 1980s by scientists at the E.R. Squibb company (now Bristol-Myers Squibb, see Note1) for their pioneering studies of C. albicans molecular biology. It was engineered by Fonzi and Irwin to provide the uridine autotrophic mutant that has been essential to most subsequent molecular genetic research into C. albicans. The strain is usually described merely as a 'clinical isolate', but it is worth setting on record that SC5314 was originally isolated from a patient with generalized Candida infection by Margarita Silva-Hutner (see Note 2) at the Department of Dermatology, Columbia College of Physicians and Surgeons (New York, USA). The original isolate number was 1775 and the strain is identical with strain NYOH#4657 in the New York State Department of Health collection. (This information was provided by Joan Fung-Tome at Bristol-Myers Squibb as a personal communication.) SC5314 belongs to the predominant clade of closely related C. albicans strains that represents almost 40% of all isolates worldwide, as determined by DNA fingerprinting and multi-locus sequence typing (A. Tavanti, A.D. Davidson, N.A.R.G., M.C.J. Maiden and F.C.O., unpublished observations)."
Note 1: The earliest publications that used SC5314 came in 1968 from Squibb Institute for Medical Research.
Note 2: The documentation of the work done by Margarita Silva-Hutner and her lab is preserved at Columbia University Archival Collections.
References: Odds et al., 2004. Genome Biol. 2004; 5(7): 230; Fonzi and Irwin, 1993. 1993 Jul;134(3):717-28; Aszalos et al., 1968. J Antibiot (Tokyo). 1968 Oct;21(10):611-5; Maestrone and Semar, 1968. Naturwissenschaften. 1968 Feb;55(2):87-8; Meyers et al., 1968. Appl Microbiol. 1968 Apr;16(4):603-8.
BWP17
Genotype: ura3::imm434/ura3::imm434 iro1/iro1::imm434 his1::hisG/his1::hisG arg4/arg4
Notes: Isogenic to the SC5314 strain. Uridine, histidine and arginine auxotroph derived from the RM1000 strain by deletion of the ARG4 gene. This strain has a heterozygous deletion on chromosome 5 that was inherited from the RM1000 parental strain.
References: Wilson et al., 1999. J Bacteriol. 1999 Mar;181(6):1868-74.
CAF2-1
Genotype: URA3/ura3::imm434 IRO1/iro1::imm434
Notes: URA3 heterozygous strain derived from the SC5314 strain. The 3-prime end of one copy of the IRO1 gene that resides adjacent to URA3 was inadvertently deleted during the construction of this strain. This strain is virulent in a mouse model of systemic infection and is frequently used as a wild-type control.
References: Fonzi and Irwin, 1993. Genetics 1993 Jul;134(3):717-28.
CAI4
Genotype: ura3::imm434/ura3::imm434 iro1/iro1::imm434
Notes: Isogenic to the SC5314 strain. Uridine auxotroph constructed by deletion of the second copy of URA3. The second copy of IRO1 was inadvertently deleted upon strain construction. As a result, the strain and its descendants have no functional copy of IRO1. This strain is avirulent in a mouse model of systemic infection unless complemented with URA3.
References: Fonzi and Irwin, 1993. Genetics 1993 Jul;134(3):717-28; Garcia et al., 2001. Yeast. 2001 Mar 15;18(4):301-11.
CAI8
Genotype: ura3::imm434/ura3::imm434 iro1/iro1::imm434 ade2::hisG/ade2::hisG
Notes: Isogenic to the SC5314 strain. Derived from the CAF2-1 strain by deletion of URA3 and both copies of ADE2 using the URA-blaster method.
References: Fonzi and Irwin, 1993. Genetics 1993 Jul;134(3):717-28.
P37005
Genotype: MTLa/MTLa
Notes: Wild-type clinical isolate. Naturally homozygous for the MTLa mating type locus.
References: Lockhart et al., 2002. Genetics. 2002 Oct;162(2):737-45.
Red3/6
Genotype: ade2/ade2
Notes: Isogenic to the WO-1 strain. Adenine auxotroph derived from the WO-1 strain by chemical mutagenesis using MNNG.
References: Srikantha et al., 1995. Mol Cell Biol. 1995 Mar;15(3):1797-805.
RM1000
Genotype: ura3::imm434/ura3::imm434 iro1/iro1::imm434 his1::hisG/his1::hisG
Notes: Isogenic to the SC5314 strain. Derived from the CAI4 strain by deletion of the HIS1 gene using the URA-blaster method (see Fonzi and Irwin, 1993 for details of this method). The standard RM1000 strain was found to have a heterozygous deletion on chromosome 5. RM1000#2 is an isolate that has been shown to have wild-type copies of chromosome 5.
References: Negredo et al., 1997. Microbiology. 1997 Feb;143 ( Pt 2):297-302.
SN87
Genotype: ura3::imm434::URA3/ura3::imm434 iro1::IRO1/iro1::imm434 his1::hisG/his1::hisG leu2/leu2
Notes: Isogenic to the SC5314 strain. Histidine and leucine auxotroph derived from the RM1000#2 strain by deletion of the LEU2 gene. This strain is virulent in a mouse model of systemic infection.
References: Noble and Johnson, 2005. Eukaryot Cell. 2005 Feb;4(2):298-309.
SN95
Genotype: ura3::imm434::URA3/ura3iro1IRO1/iro1his1his1arg4/arg4
Notes: Isogenic to the SC5314 strain. Histidine and arginine auxotroph derived from the RM1000#2 strain by deletion of the ARG4 gene. This strain is virulent in a mouse model of systemic infection.
References: Noble and Johnson, 2005. Eukaryot Cell. 2005 Feb;4(2):298-309.
SN152
Genotype: ura3/::imm434::URA3/ura3::imm434 iro1::IRO1/iro1::imm434 his1::hisG/his1::hisG leu2/leu2 arg4/arg4
Notes: Isogenic to the SC5314 strain. Histidine, leucine and arginine auxotroph derived from the RM1000#2 strain by deletion of the LEU2 and ARG4 genes. This strain is virulent in a mouse model of systemic infection.
References: Noble and Johnson, 2005. Eukaryot Cell. 2005 Feb;4(2):298-309.
SN250
Genotype: his1Δ/his1Δ, leu2Δ::C.dubliniensis HIS1 /leu2Δ::C.maltosa LEU2, arg4Δ /arg4Δ, URA3/ura3Δ::imm434, IRO1/iro1Δ::imm434
Notes: Isogenic to the SC5314 strain. Derived from SN87 by integration of C. dubliniensis HIS1 and C. maltosa LEU2 at the disrupted leu2 loci and then deleted for arg4. Derived from SN87 and QMY23.
References: Noble et al., 2010. Nat Genet. 2010 Jul;42(7):590-8;
Mitrovich et al., 2007. Genome Res. 2007 Apr;17(4):492-502.
WO-1
Genotype: MTLalpha
Notes: Wild-type clinical isolate that switches between white and opaque phenotypes at high frequency. The MTLa locus is absent in this strain (for more information see Lockhart et al., 2002). This strain has been sequenced by the Broad Institute (http://www.broadinstitute.org/annotation/genome/candida_group/GenomeDescriptions.html)
References: Slutsky et al., 1987. J Bacteriol. 1987 Jan;169(1):189-97; Lockhart et al., 2002. Genetics. 2002 Oct;162(2):737-45.
WUM5A
Genotype: MTLalpha/MTLalpha ura3-1Δ::FRT/ura3-2Δ::FRT
Notes: Isogenic to the WO-1 strain. Uridine auxotroph constructed by deletion of both copies of URA3.
References: Strauss et al. 2001. J Bacteriol. 2001 Jun;183(12):3761-9.
Candida glabrata (Nakaseomyces glabratus) Strains
B
Genotype: Wild type
Notes: Clinical isolate from a case of vaginitis that did not respond to fluconazole or boric acid treatment; it is virulent in a murine model of vaginitis. This strain was called LF 547.92 in the original publication.
References: Fidel et al., 1996. J Infect Dis. 1996 Feb;173(2):425-31.
BG2
Genotype: Wild type
Notes: Clinical isolate from a case of vaginitis that did not respond to fluconazole or boric acid treatment; it is virulent in a murine model of vaginitis
References: Cormack and Falkow, 1999. Genetics. 1999 Mar;151(3):979-87.
BG14
Genotype: ura3-delta(-285 +932)::Tn903NeoR
Notes: A ura3 derivative of the wild-type B2 clinical isolate.
References: Cormack and Falkow, 1999. Genetics. 1999 Mar;151(3):979-87.
BG462
Genotype: URA3
Notes: A derivative of the B2 clinical isolate with the URA3 gene restored
References: De Las Peñas, et al., 2003. Genes Dev. 2003 Sep 15;17(18):2245-58.
CBS138 (ATCC 2001)
Genotype: Wild type
Notes: Type strain for the Genolevures C. glabrata sequencing project. This strain has also been called NRRL-Y-65 and IFO 0622.
References: Dujon et al., 2004. Nature. 2004 Jul 1;430(6995):35-44; Koszul, et al., 2003. FEBS Lett. 2003 Jan 16;534(1-3):39-48.
2001T
Genotype: trp1
Notes: A trp1 auxotrophic strain derived from the CBS138 (ATCC 2001) wild-type strain.
References: Kitada et al., 1995. Gene. 1995 Nov 20;165(2):203-6.
2001HT
Genotype: his3 trp1::ScURA3
Notes: A his3 trp1 auxotrophic strain derived from 200T, a derivative of the CBS138 (ATCC 2001) wild-type strain.
References: Kitada et al., 1995. Gene. 1995 Nov 20;165(2):203-6.
NCCLS84 (ATCC 90030)
Genotype: Wild type
Notes: Wild-type strain
References: Espinel-Ingroff et al., 1992. J Clin Microbiol. 1992 Dec;30(12):3138-45
84u
Genotype: ura3
Notes: A ura3 derivative of the wild-type NCCLS84 (ATCC90030) strain.
References: Tsai et al., 2006. Antimicrob Agents Chemother. 2006 Apr;50(4):1384-92.
Candida parapsilosis Strains
ATCC 22019
Genotype: heterozygous MET/met
Notes: Application of parasexual genetic methods was used to determine that C. parapsilosis strain ATCC 22019 is is heterozygous MET/met
References: Whelan and Kwon-Chung, 1988. J Med Vet Mycol 1988 Jun;26(3):163-71.
CDC317
Genotype: Wild type
Notes: Reference strain used for the C. parapsilosis sequencing project. This isolate came from the hands of a hospital worker, who was the source for an outbreak of infection in a Mississippi community hospital in 2001.
References: Kuhn et al., 2004. Emerg Infect Dis. 2004 Jun;10(6):1074-81; Clark et al., 2004. J Clin Microbiol. 2004 Oct;42(10):4468-72.
CLIB214 (CBS 604; ATCC 22019; NRRL Y-12969)
Genotype: Wild type
Notes: Type strain, originally isolated from a patient with diarrhea in Puerto Rico by Ashford (1928). This strain is dependent on oxidative metabolism for growth since it lacks a fermentative pathway.
References: Ashford, 1928. Am J Trop Med 8:507�538. In: Kurtzman CP, Fell JW, Boekhout T (eds), 2011. The yeasts: a taxonomic study. p. 987-1278, 5th ed. Elsevier, London, United Kingdom; Logue et al., 2005. Eukaryot Cell 4(6):1009-17.
CPL2
Genotype: leu2Δ::FRT/leu2Δ::FRT, his1Δ::FRT/his1Δ::FRT, frt::CdHIS1
Notes: Leucine auxotroph laboratory strain
References: Németh et al., 2021. Virulence. 2021 Dec;12(1):937-950.
GA1
Genotype: Wild type
Notes: Wild-type clinical isolate. The SAT1 flipper method and the Candida albicans IMH3 gene have been used as dominant-selectable markers to construct gene knockouts in this strain.
References: Gácser et al., 2005. FEMS Microbiol Lett. 2005 Apr 1;245(1):117-21; Gácser et al., 2007. J Clin Invest. 2007 Oct;117(10):3049-58.
SR23 (CBS7157)
Genotype: ade- lys-
Notes: An adenine and lysine auxotroph isolated from a contaminated culture of Saccharomyces cerevisiae as a yeast with peculiar physiological features and carrying a linear mitochondrial DNA
References: Kovac et al., 1984. Mol Gen Genet 197:420-424; Nosek et al., 2004. Mol Genet Genomics 272(2):173-80; Mutalová et al., 2024. Microbiol Resour Announc. 2024 Jul 31;13(9):e00347-24.
Candida auris (Candidozyma auris) Strains
AR0382 (B11109)
Genotype: Wild type
Notes: Aggregative strain with high biofilm formation.
References: Vila et al., 2020. mSphere. 2020 Aug 5;5(4):e00760-20; Wang et al., 2024. Nat Commun. 2024 Oct 25;15(1):9212.
AR0387 (B8441)
Genotype: Wild type
Notes: Reference strain; non-aggregative strain with low biofilm formation.
References: Vila et al., 2020. mSphere. 2020 Aug 5;5(4):e00760-20; Wang et al., 2024. Nat Commun. 2024 Oct 25;15(1):9212.
AR0389
Genotype: erg11-Y132F
Notes: Has high drug resistance due to an ERG11 mutation (Y132F) and overexpression of the drug efflux transporter gene CDR1.
References: Shinohara et al., 2024. JAC Antimicrob Resist. 2024 Feb 21;6(1):dlad155.
AR0390 (B11205)
Genotype: erg11-K143R
Notes: Clinical isolate with high fluconazole resistance.
References: Rybak et al., 2021. Microbiol Spectr. 2021 Dec 8;9(3):e01585-21.
B11220
Genotype: Wild type
Notes: Clade II clinical isolate with susceptibility to all antifungal drugs.
References: Yang et al., 2024. Infect Immun. 2024 Jun 11;92(6):e0010324.
B11221
Genotype: Wild type
Notes: Clade III clinical isolate with resistance to caspofungin. Contains an L-rhamnose utilization cluster that appears to allow more ready use of alternate sugars, better drug tolerance, and better survival from host immune cell attack, possibly due to reduced β-(1,3)-glucans at the cell surface.
References: Yang et al., 2024. Infect Immun. 2024 Jun 11;92(6):e0010324.
Candida tropicalis Strains
ATCC 20336 (pK233)
Genotype: Wild type
Notes: This strain excretes alpha,omega-dicarboxylic acids as a by-product when cultured on n-alkanes or fatty acids as the carbon source.
References: Craft et al., 2003. Appl Environ Microbiol 69(10):5983-91.
ATCC 20913
Genotype: ura3/ura3
Notes: A uracil auxotroph derived from C. tropicalis ATCC 20336 by random mutagenesis.
References: Haas et al., 1990. J Bacteriol 172(8):4571-7.
ATCC 750
Genotype: Wild type
Notes: A wild-type fluconazole susceptible strain.
References: Barchiesi et al., 2000. Antimicrob Agents Chemother 44(6):1578-84.
NCYC2512
Genotype: Wild type
Notes: A wild-type saline soil isolate from Pakistan that is capable of producing large amounts of alpha,omega-dodecanedioic acid.
References: Rodriguez et al., 1996. Yeast 12(13):1321-9.
1230
Genotype: Wild type
Notes: A wild-type dicarboxylic acid-producing industrial strain.
References: He and Chen, 2005. Yeast 22(6):481-91.
Candida dubliniensis Strains
CD36
Genotype: Wild type
Notes: Type strain, oral isolate from an HIV+ patient in Ireland; also catalogued as NCPF 3949 and CBS-7987.
References: Sullivan et al., 1995. Microbiology (Reading) 1995 Jul:141 ( Pt 7):1507-21.
Protocols and Methods
Candida albicans Protocols and Methods
- Basic techniques
- Candida albicans: Methods and Protocols
- Cihlar and Calderone, 2009. Methods Mol Biol 2009:499:v
- Insertion mutagenesis
- Mielich et al., 2018 G3 (Bethesda) 2018 Mar 28;8(4):1139-1145
- CRISPR-Cas9 deletions
- Halder et al., 2019. Nat Protoc. 2019 Mar;14(3):955-975
- Shapiro et al., 2018. Nat Microbiol. 2018 Jan;3(1):73-82
- Wensing and Shapiro, 2022. Methods Mol Biol. 2022:2377:69-88
- Nguyen et al., 2021. Microbiol Spectr. 2021 Dec 22;9(3):e0182021
- 5-FOA assay for assessing mutation rates
- Lang and Murray, 2008. Genetics. 2008 Jan;178(1):67-82
- Virulence studies in Galleria mellonella
- Champion et al., 2016. Virulence. 2016 Oct 2;7(7):840-5
- Desbois and Coote, 2016. Virulence. 2016 Oct 2;7(7):840-5
- Junqueira, 2012. Virulence. 2012 Oct 1;3(6):474-6
- Wojda et al., 2020. Pathog Dis. 2020 Nov 23;78(9):ftaa057
Candida glabrata (Nakaseomyces glabratus) Protocols and Methods
- Basic techniques
- MIC assays
- Clinical and Laboratory Standards Performance Standards for Antifungal Susceptibility Testing of Yeasts, 3rd Edition
- Israel et al., 2021. J Clin Microbiol. 2021 Jun 18;59(7):e00311-21
- Genetic manipulation of C. glabrata
- Usher, 2024. Curr Protoc. 2024 Sep;4(9):e70014
- Transformation of C. glabrata
- Istel et al., 2015. Bio Protoc. 2015 Jul 20;5(14):e1528
- Istel et al., 2015. Bio Protoc. 2015 Jul 20;5(14):e1529
- CRISPR-Cas9-based deletions
- Enkler et al., 2016. Sci Rep. 2016 Oct 21:6:35766
- Maroc and Fairhead, 2019. Yeast. 2019 Dec;36(12):723-731
- Virulence studies in Galleria mellonella
- Champion et al., 2016. Virulence. 2016 Oct 2;7(7):840-5
- Desbois and Coote, 2016. Virulence. 2016 Oct 2;7(7):840-5
- Junqueira, 2012. Virulence. 2012 Oct 1;3(6):474-6
- Wojda et al., 2020. Pathog Dis. 2020 Nov 23;78(9):ftaa057
Candida parapsilosis Protocols and Methods
- Basic techniques
- Candida Species: Methods and Protocols
- Arévalo-Jaimes et al.,2023. Front Cell Infect Microbiol. 2023 Dec 13;13:1323619.
- Transformation of C. parapsilosis
- Nemeth et al.,2021. Virulence. 2021 Mar 17;12(1):937–950
- Zemanova et al.,2004. Curr Genet. 2004 Mar;45(3):183-6
- Virulence studies in Galleria mellonella
- Champion et al., 2016. Virulence. 2016 Oct 2;7(7):840-5
- Desbois and Coote, 2016. Virulence. 2016 Oct 2;7(7):840-5
- Junqueira, 2012. Virulence. 2012 Oct 1;3(6):474-6
- Wojda et al., 2020. Pathog Dis. 2020 Nov 23;78(9):ftaa057
- CRISPR-Cas9-based deletions
- Lombardi et al.,2022. mSphere. 2022 Dec 21;7(6):e0039322
- Lombardi and Butler,2022. Methods Mol Biol. 2022:2542:13-40
Candida auris (Candidozyma auris) Protocols and Methods
- Basic techniques
- Identification of C. auris
- Narayanan et al., 2022. Microbiol Spectr. 2022 Apr 27;10(2):e0063422
- Abdolrasouli and Fraser, 2022. Methods Mol Biol. 2022:2517:21-32
- Transformation of C. auris
- Ruiz and Lorenz, 2022. Methods Mol Biol. 2022:2517:95-110
- Kasir et al., 2022. Methods Mol Biol. 2022: 2517:89-94
- CRISPR-Cas9-based deletions
- Ennis et al., 2021. Microbiol Spectr. 2021 Dec 22;9(3):e0182021
- Nguyen et al., 2017. mSphere. 2017 Apr 26;2(2):e00149-17.
- Virulence studies in Galleria mellonella
- Borman et al., 2016. mSphere. 2016 Aug 18;1(4):e00189-16
- Garcia-Bustos et al., 2022. Emerg Microbes Infect. 2022 Dec;11(1):136-146
Candida dubliensis Protocols and Methods
- Basic techniques
- Identification
- Tintelnot at al., 2000. J Clin Microbiol. 2000 Apr;38(4):1599-608
- Bliss et al., 2003. J Clin Microbiol. 2003 Mar;41(3):1152-60
- Ells et al., 2011. Mycoses. 2011 Jan;54(1):1-16
- Kianipour et al., 2018. Adv Biomed Res. 2018 Apr 24:7:66
- Transformation and strain construction in C. dubliniensis
- Staib et al., 2000. Gene. 2000 Jan 25;242(1-2):393-8
- Staib et al., 2001. J Bacteriol. 2001 May;183(9):2859-65
- Virulence studies in Galleria mellonella
- Chupácová et al., 2018. Pathog Dis. 2018 Feb 1;76(1)
- Dekkerová-Chupáčová et al., 2018. Mycopathologia. 2018 Dec;183(6)
Candida tropicalis Protocols and Methods
- Basic techniques
- Transformation of C. tropicalis
- Rohrer and Picataggio, 1992. Appl Microbiol Biotechnol. 1992 Feb;36(5):650-4
- Xiang et al., 2014. Yi Chuan. 2014 Oct;36(10):1053-61
- CRISPR-Cas9-based deletions
- Li et al., 2022. Microbiol Spectr. 2022 Jun 29;10(3):e0005922
- Lombardi and Butler, 2022. Methods Mol Biol. 2022:2542:13-40
- Zhang et al., 2020. Biotechnol Bioeng. 2020 Feb;117(2):531-542
- Virulence studies in Galleria mellonella
- Mesa-Arango et al., 2013. Med Mycol. 2013 Jul;51(5):461-72
- Forastiero et al., 2013. Antimicrob Agents Chemother. 2013 Oct;57(10):4769-81
- García-Carnero et al., 2020. J Fungi (Basel). 2020 Aug 27;6(3):152
Species Comparisons by Topic
Ploidy
Notes: Candida albicans and its close relatives are all on a single phylogenetic branch characterized by diploidy, while the two typical haploid species (C. glabrata and C. auris) are on distinct, distantly related branches for which haploidy is the standard form. Note that the two haploid pathogens are more prone to pleiotropic resistance because only a single mutation is needed in the haploid genome to generate stable change.
References:
Typical diploids
- C. albicans
- C. dubliniensis
- C. parapsilosis
- C. tropicalis
Typical haploids
- C. glabrata
- C. auris
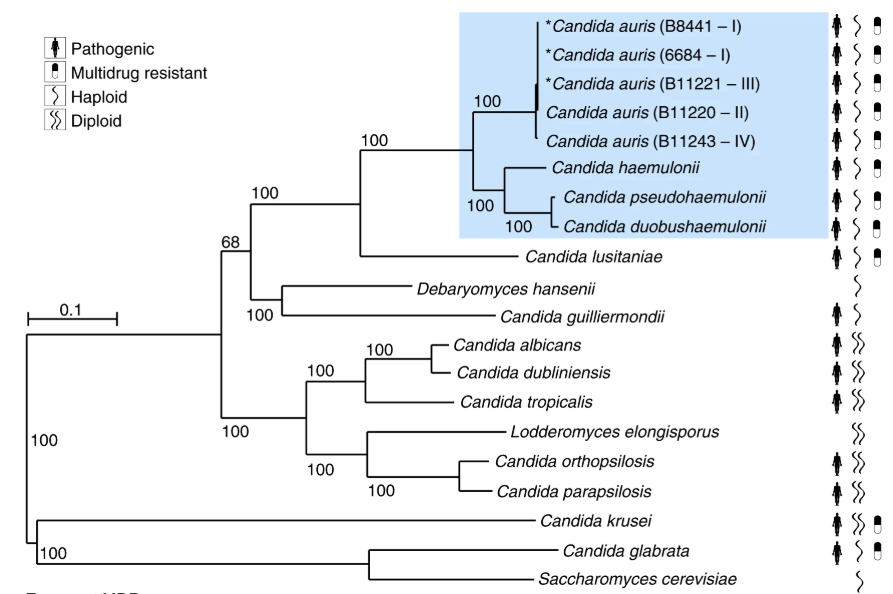
From Munoz et al., 2018
CUG codon usage
Notes: Yeasts of the Candida clade (as opposed to C. glabrata, also known as Nakaseomyces glabratus, which is on the Saccharomyces clade) have undergone an evolutionary reassignment of the CUG codon from leucine to serine. This is due to a novel serine-tRNA (ser-tRNACAG) that contains a guanosine at position 33, a position that is occupied by a pyrimidine (uridine) in nearly all other tRNAs.
References:
Reassigned CUG as serine
- C. albicans
- C. dubliniensis
- C. parapsilosis
- C. tropicalis
- C. auris
Standard CUG as leucine
- C. glabrata
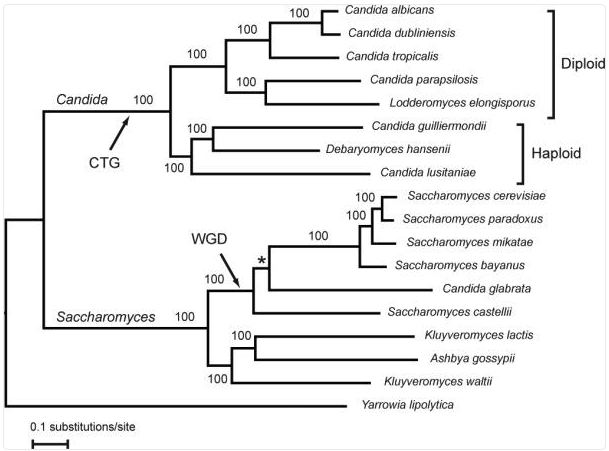
From Butler et al., 2009
Mitochondrial genome
Notes: Candida parapsilosis is unusual in having a mitochondrial genome that is linear and capped by telomeric ends. The close relatives of this species, C. orthopsilosis and C. metapsilosis and Candida subhashii, have a mix of circular and linear mitochondrial genomes. The other Candida spp. are exclusively circular.
References:
Exclusively linear
- C. parapsilosis
Mix of linear and circular
- C. orthopsilosis
- C. metapsilosis
- C. subhashii
Exclusively circular
- C. albicans
- C. dubliniensis
- C. tropicalis
- C. auris
- C. glabrata
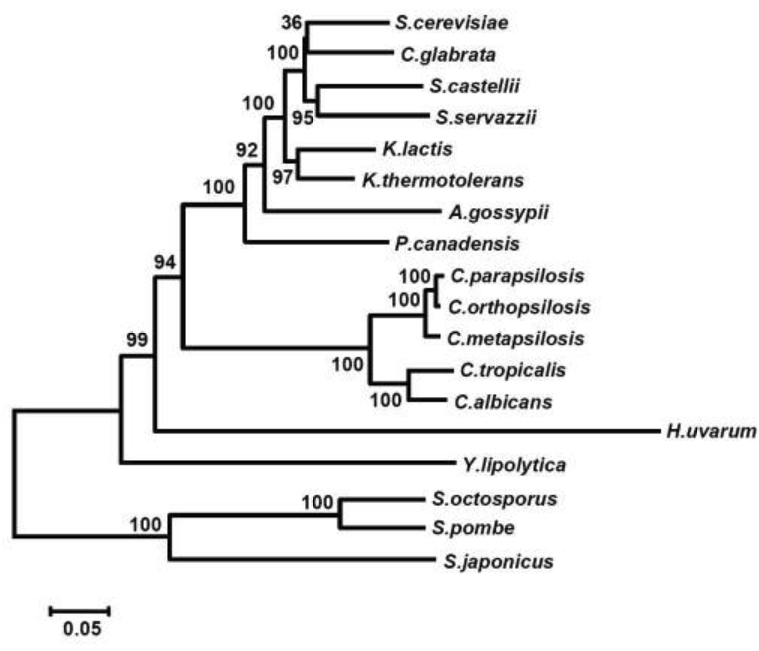
From Kosa et al., 2006
Filamentous growth
Notes: Candida spp. vary in filamentous growth as a means of infection/invasion. C. albicans and its close relative C. dubliniensis use true hyphae to penetrate tissues, while C. parapsilosis and C. tropicalis can penetrate tissues by means of pseudohyphae (lacking septa). In contrast, C. glabrata almost never forms filaments and infects in the yeast form. C. auris is an outlier in that it infects both by pseudohyphae and in the yeast form.
Note that the classifications below represent the typical forms for each species, but exceptions are not unusual.
References:
- Arita et al., 2022 (review)
- Kornitzer, 2019 (review)
- Silva et al., 2012 (review)
- Branco et al., 2023 (review)
- Bryak et al., 2024
- Groenewald et al., 2023
Invasive/infectious by true hyphae
- C. albicans
- C. dubliniensis
Invasive/infectious pseudohyphal growth (lacking septa)
- C. parapsilosis
- C. tropicalis
- C. auris
Invasive/infectious as yeast form
- C. glabrata
- C. auris

From Kosa et al., 2006
Biofilms
Pathogenicity
Mating/Reproduction
Drug resistance
Respiratory metabolism
Datasets
- RNA-Seq
- ChIP-Seq