Main Page
Welcome to the CGD Public Wiki
Seminal Candida Papers
C. albicans
Cell Biology
AUTOPHAGY
- Palmer GE, Kelly MN, Sturtevant JE. Autophagy in the pathogen Candida albicans. Microbiology. 2007 Jan;153(Pt 1):51-8.
CELL CYCLE
- Barton R, Gull K. Variation in cytoplasmic microtubule organization and spindle length between the two forms of the dimorphic fungus Candida albicans. J Cell Sci. 1988 Oct;91 ( Pt 2):211-20.
- Berman, J. Morphogenesis and cell cycle progression in Candida albicans. 2006 Dec;9(6):595-601. Epub 2006 Oct 20.
- Chapa y Lazo B, Bates S, Sudbery P. The G1 cyclin Cln3 regulates morphogenesis in Candida albicans. Eukaryot Cell. 2005 Jan;4(1):90-4.
- Cote P, Hogues H, Whiteway M. Transcriptional analysis of the Candida albicans cell cycle. Mol Biol Cell. 2009 Jul;20(14):3363-73. Epub 2009 May 28.
- Finley KR, Berman J. Microtubules in Candida albicans hyphae drive nuclear dynamics and connect cell cycle progression to morphogenesis. Eukaryot Cell. 2005 Oct;4(10):1697-711.
- Singh A, Sharma S, Khuller GK. cAMP regulates vegetative growth and cell cycle in Candida albicans. Mol Cell Biochem. 2007 Oct;304(1-2):331-41. Epub 2007 Jun 8.
CELL WALL
- Chaffin WL. Candida albicans cell wall proteins. Microbiol Mol Biol Rev. 2008 Sep;72(3):495-544.
- Ebanks RO, Chisholm K, McKinnon S, Whiteway M, Pinto DM. Proteomic analysis of Candida albicans yeast and hyphal cell wall and associated proteins. Proteomics. 2006 Apr;6(7):2147-56.
- Klis FM, de Groot P, Hellingwerf K. Molecular organization of the cell wall of Candida albicans. Med Mycol. 2001;39 Suppl 1:1-8.
- Klis FM, Sosinska GJ, de Groot PW, Brul S. Covalently linked cell wall proteins of Candida albicans and their role in fitness and virulence. FEMS Yeast Res. 2009 Oct;9(7):1013-28. Epub 2009 Jun 22.
- Ruiz-Herrera J, Elorza MV, Valentin E, Sentandreu R. Molecular organization of the cell wall of Candida albicans and its relation to pathogenicity. FEMS Yeast Res. 2006 Jan;6(1):14-29.
- Sohn K, Schwenk J, Urban C, Lechner J, Schweikert M, Rupp S. Getting in touch with Candida albicans: the cell wall of a fungal pathogen. Curr Drug Targets. 2006 Apr;7(4):505-12.
- Sundstrom P. Adhesion in Candida spp. Cell Microbiol. 2002 Aug;4(8):461-9.
- Gow NA, Hube B. Importance of the Candida albicans cell wall during commensalism and infection. Curr Opin Microbiol. 2012 Aug;15(4):406-12.
- Gow NAR, Latge JP, Munro CA. The fungal cell wall: structure, biosynthesis, and function. Microbiol Spectr. 2017 May;5(3)
- Gow NAR, Lenardon MD. Architecture of the dynamic fungal cell wall. Nat Rev Microbiol. 2023 Apr;21(4):248-259.
CO2 SENSING
- Mitchell AP. Fungal CO2 sensing: a breath of fresh air. Curr Biol. 2005 Nov 22;15(22):R934-6.
- Mock RC, Pollack JH, Hashimoto T. Carbon dioxide induces endotrophic germ tube formation in Candida albicans. Can J Microbiol. 1990 Apr;36(4):249-53.
- Sims W. Effect of carbon dioxide on the growth and form of Candida albicans. J Med Microbiol. 1986 Nov;22(3):203-8.
- Webster CE, Odds FC. Growth of pathogenic Candida isolates anaerobically and under elevated concentrations of CO2 in air. J Med Vet Mycol. 1987 Feb;25(1):47-53. Erratum in: J Med Vet Mycol 1988 Feb;26(1):75.
GENETIC INSTABILITY
- Selmecki A, Bergmann S, Berman J. Comparative genome hybridization reveals widespread aneuploidy in Candida albicans laboratory strains. Mol Microbiol. 2005 Mar;55(5):1553-65.
- Selmecki AM, Dulmage K, Cowen LE, Anderson JB, Berman J. Acquisition of aneuploidy provides increased fitness during the evolution of antifungal drug resistance. PLoS Genet. 2009 Oct;5(10):e1000705. Epub 2009 Oct 30.
- Wu W, Pujol C, Lockhart SR, Soll DR. Chromosome loss followed by duplication is the major mechanism of spontaneous mating-type locus homozygosis in Candida albicans. Genetics. 2005 Mar;169(3):1311-27. Epub 2005 Jan 16.
- Berman J. Ploidy plasticity: a rapid and reversible strategy for adaptation to stress. FEMS Yeast Res. 2016 May;16(3):fow020.
VACUOLAR DYNAMICS AND INHERITANCE
- Gow NA, Gooday GW. Growth kinetics and morphology of colonies of the filamentous form of Candida albicans. J Gen Microbiol. 1982 Sep;128(9):2187-94.
- Gow NA, Gooday GW. A model for the germ tube formation and mycelial growth form of Candida albicans. Sabouraudia. 1984;22(2):137-44.
- Barelle CJ, Bohula EA, Kron SJ, Wessels D, Soll DR, Schafer A, Brown AJ, Gow NA. Asynchronous cell cycle and asymmetric vacuolar inheritance in true hyphae of Candida albicans. Eukaryot Cell. 2003 Jun;2(3):398-410.
- Veses V, Gow NA. Vacuolar dynamics during the morphogenetic transition in Candida albicans. FEMS Yeast Res. 2008 Dec;8(8):1339-48.
- Veses V, Richards A, Gow NA. Vacuole inheritance regulates cell size and branching frequency of Candida albicans hyphae. Mol Microbiol. 2009 Jan;71(2):505-19.
Transformation/Response
BIOFILMS
- Dongari-Bagtzoglou A, Kashleva H, Dwivedi P, Diaz P, Vasilakos J. Characterization of mucosal Candida albicans biofilms. PLoS One. 2009 Nov 24;4(11):e7967.
- Ene IV, Bennett RJ. Hwp1 and related adhesins contribute to both mating and biofilm formation in Candida albicans. Eukaryot Cell. 2009 Dec;8(12):1909-13. Epub 2009 Oct 16
- Kumamoto CA, Vinces MD. Alternative Candida albicans lifestyles: growth on surfaces. Annu Rev Microbiol. 2005;59:113-33.
- Murillo LA, Newport G, Lan CY, Habelitz S, Dungan J, Agabian NM. Genome-wide transcription profiling of the early phase of biofilm formation by Candida albicans. Eukaryot Cell. 2005 Sep;4(9):1562-73.
- Nobile CJ, Schneider HA, Nett JE, Sheppard DC, Filler SG, Andes DR, Mitchell AP. Complementary adhesin function in C. albicans biofilm formation. Curr Biol. 2008 Jul 22;18(14):1017-24.
- Thomas DP, Bachmann SP, Lopez-Ribot JL. Proteomics for the analysis of the Candida albicans biofilm lifestyle. Proteomics. 2006 Nov;6(21):5795-804.
- ten Cate JM, Klis FM, Pereira-Cenci T, Crielaard W, de Groot PW. Molecular and cellular mechanisms that lead to Candida biofilm formation. J Dent Res. 2009 Feb;88(2):105-15.
- Soll DR, Daniels KJ. Plasticity of Candida albicans Biofilms. Microbiol Mol Biol Rev. 2016 Jun 1;80(3):565-95.
- Nobile CJ, Mitchell AP. Genetics and genomics of Candida albicans biofilm formation. Cell Microbiol. 2006 Sep;8(9):1382-91.
- Gulati M, Nobile CJ. Candida albicans biofilms: development, regulation, and molecular mechanisms. Microbes Infect. 2016 May;18(5):310-21.
- Lohse MB, Gulati M, Johnson AD, Nobile CJ. Development and regulation of single- and multi-species Candida albicans biofilms.
Nat Rev Microbiol. 2018 Jan;16(1):19-31.
- Nobile CJ, Johnson AD. Candida albicans biofilms and human disease. Annu Rev Microbiol. 2015;69:71-92.
- Perry AM, Hernday AD, Nobile CJ. Unraveling How Candida albicans Forms Sexual Biofilms. J Fungi (Basel). 2020 Jan 15;6(1):14.
CHLAMYDOSPORE DEVELOPMENT
- [https://pubmed.ncbi.nlm.nih.gov/12627804/ Fabry W, Schmid EN, Schraps M, Ansorg R. Isolation and purification of chlamydospores of Candida albicans. Med Mycol. 2003 Feb;41(1):53-8.
- Jansons VK, Nickerson WJ. Chemical composition of chlamydospores of Candida albicans. J Bacteriol. 1970 Nov;104(2):922-32.
- Martin SW, Douglas LM, Konopka JB. Cell cycle dynamics and quorum sensing in Candida albicans chlamydospores are distinct from budding and hyphal growth. Eukaryot Cell. 2005 Jul;4(7):1191-202.
- Nobile CJ, Bruno VM, Richard ML, Davis DA, Mitchell AP. Genetic control of chlamydospore formation in Candida albicans. Microbiology. 2003 Dec;149(Pt 12):3629-37.
HYPOXIA
- Ernst JF, Tielker D. Responses to hypoxia in fungal pathogens. Cell Microbiol. 2009 Feb;11(2):183-90.
- Mulhern SM, Logue ME, Butler G. Candida albicans transcription factor Ace2 regulates metabolism and is required for filamentation in hypoxic conditions. Eukaryot Cell. 2006 Dec;5(12):2001-13.
- Setiadi ER, Doedt T, Cottier F, Noffz C, Ernst JF. Transcriptional response of Candida albicans to hypoxia: linkage of oxygen sensing and Efg1p-regulatory networks. J Mol Biol. 2006 Aug 18;361(3):399-411.
- Stichternoth C, Ernst JF. Hypoxic adaptation by Efg1 regulates biofilm formation by Candida albicans. Appl Environ Microbiol. 2009 Jun;75(11):3663-72.
MATING AND THE PARASEXUAL CYCLE
- Hull CM, Johnson AD. Identification of a mating type-like locus in the asexual pathogenic yeast Candida albicans. Science. 1999 Aug 20;285(5431):1271-5.
- Hull CM, Raisner RM, Johnson AD. Evidence for mating of the "asexual" yeast Candida albicans in a mammalian host. Science. 2000 Jul 14;289(5477):307-10.
- Magee BB, Magee PT. Induction of mating in Candida albicans by construction of MTLa and MTLalpha strains. Science. 2000 Jul 14;289(5477):310-3.
- Lockhart SR, Pujol C, Daniels KJ, Miller MG, Johnson AD, Pfaller MA, Soll DR. In Candida albicans, white-opaque switchers are homozygous for mating type. Genetics. 2002 Oct;162(2):737-45.
- Alby K, Schaefer D, Bennett RJ. Homothallic and heterothallic mating in the opportunistic pathogen Candida albicans. Nature. 2009 Aug 13;460(7257):890-3.
- Bennett RJ, Johnson AD. Completion of a parasexual cycle in Candida albicans by induced chromosome loss in tetraploid strains. EMBO J. 2003 May 15;22(10):2505-15.
- Bennett RJ, Uhl MA, Miller MG, Johnson AD. Identification and characterization of a Candida albicans mating pheromone. Mol Cell Biol. 2003 Nov;23(22):8189-201.
- Cote P, Whiteway M. The role of Candida albicans FAR1 in regulation of pheromone-mediated mating, gene expression and cell cycle arrest. Mol Microbiol. 2008 Apr;68(2):392-404.
- Daniels KJ, Lockhart SR, Staab JF, Sundstrom P, Soll DR. The adhesin Hwp1 and the first daughter cell localize to the a/a portion of the conjugation bridge during Candida albicans mating. Mol Biol Cell. 2003 Dec;14(12):4920-30.
- Ene IV, Bennett RJ. Hwp1 and related adhesins contribute to both mating and biofilm formation in Candida albicans. Eukaryot Cell. 2009 Dec;8(12):1909-13.
- Forche A, Alby K, Schaefer D, Johnson AD, Berman J, Bennett RJ. The parasexual cycle in Candida albicans provides an alternative pathway to meiosis for the formation of recombinant strains. PLoS Biol. 2008 May 6;6(5):e110.
- Ibrahim AS, Magee BB, Sheppard DC, Yang M, Kauffman S, Becker J, Edwards JE Jr, Magee PT. Effects of ploidy and mating type on virulence of Candida albicans. Infect Immun. 2005 Nov;73(11):7366-74.
- Lockhart SR, Daniels KJ, Zhao R, Wessels D, Soll DR. Cell biology of mating in Candida albicans. Eukaryot Cell. 2003 Feb;2(1):49-61.
- Lockhart SR, Wu W, Radke JB, Zhao R, Soll DR. Increased virulence and competitive advantage of a/alpha over a/a or alpha/alpha offspring conserves the mating system of Candida albicans. Genetics. 2005 Apr;169(4):1883-90.
- Lockhart SR, Zhao R, Daniels KJ, Soll DR. Alpha-pheromone-induced "shmooing" and gene regulation require white-opaque switching during Candida albicans mating. Eukaryot Cell. 2003 Oct;2(5):847-55.
- Sherwood RK, Bennett RJ. Fungal meiosis and parasexual reproduction--lessons from pathogenic yeast. Curr Opin Microbiol. 2009 Dec;12(6):599-607.
- Soll DR, Lockhart SR, Zhao R. Relationship between switching and mating in Candida albicans. Eukaryot Cell. 2003 Jun;2(3):390-7.
- Wu W, Pujol C, Lockhart SR, Soll DR. Chromosome loss followed by duplication is the major mechanism of spontaneous mating-type locus homozygosis in Candida albicans. Genetics. 2005 Mar;169(3):1311-27.
- Bennett RJ, Johnson AD. Mating in Candida albicans and the search for a sexual cycle. Annu Rev Microbiol. 2005;59:233-55.
- Wolfe KH, Butler G. Evolution of Mating in the Saccharomycotina. Annu Rev Microbiol. 2017 Sep 8;71:197-214.
MORPHOGENESIS AND POLARIZED CELL GROWTH
- Gow NA, Gooday GW. Growth kinetics and morphology of colonies of the filamentous form of Candida albicans. J Gen Microbiol. 1982 Sep;128(9):2187-94.
- Gow NA, Gooday GW. Vacuolation, branch production and linear growth of germ tubes in Candida albicans. J Gen Microbiol. 1982 Sep;128(9):2195-8.
- [https://pubmed.ncbi.nlm.nih.gov/3319423/ Gow NA, Gooday GW. Cytological aspects of dimorphism in Candida albicans. Crit Rev Microbiol. 1987;15(1):73-8.
- Gow NA. Germ tube growth of Candida albicans. Curr Top Med Mycol. 1997 Dec;8(1-2):43-55.
- Berman, J. Morphogenesis and cell cycle progression in Candida albicans. 2006 Dec;9(6):595-601.
- Sinha I, Wang YM, Philp R, Li CR, Yap WH, Wang Y. Cyclin-dependent kinases control septin phosphorylation in Candida albicans hyphal development. Dev Cell. 2007 Sep;13(3):421-32.
- Veses V, Gow NA. Vacuolar dynamics during the morphogenetic transition in Candida albicans. FEMS Yeast Res. 2008 Dec;8(8):1339-48.
- Veses V, Richards A, Gow NA. Vacuole inheritance regulates cell size and branching frequency of Candida albicans hyphae. Mol Microbiol. 2009 Jan;71(2):505-19.
- Wang Y. CDKs and the yeast-hyphal decision. Curr Opin Microbiol. 2009 Dec;12(6):644-9.
- Whiteway M, Bachewich C. Morphogenesis in Candida albicans. Annu Rev Microbiol. 2007;61:529-53.
- Du H, Ennis CL, Hernday AD, Nobile CJ, Huang G. N-Acetylglucosamine (GlcNAc) Sensing, Utilization, and Functions in Candida albicans. J Fungi (Basel). 2020 Aug 7;6(3):129.
WHITE-OPAQUE SWITCHING
- Anderson J, Cundiff L, Schnars B, Gao MX, Mackenzie I, Soll DR. Hypha formation in the white-opaque transition of Candida albicans. Infect Immun. 1989 Feb;57(2):458-67.
- Anderson J, Mihalik R, Soll DR. Ultrastructure and antigenicity of the unique cell wall pimple of the Candida opaque phenotype. J Bacteriol. 1990 Jan;172(1):224-35.
- Soll DR. High-frequency switching in Candida albicans. Clin Microbiol Rev. 1992 Apr;5(2):183-203.
- Soll DR, Morrow B, Srikantha T. High-frequency phenotypic switching in Candida albicans. Trends Genet. 1993 Feb;9(2):61-5.
- Soll DR, Lockhart SR, Zhao R. Relationship between switching and mating in Candida albicans. Eukaryot Cell. 2003 Jun;2(3):390-7.
- Lohse MB, Johnson AD. White-opaque switching in Candida albicans. Curr Opin Microbiol. 2009 Dec;12(6):650-4.
- Alby K, Bennett RJ. Stress-induced phenotypic switching in Candida albicans. Mol Biol Cell. 2009 Jul;20(14):3178-91.
- Daniels KJ, Srikantha T, Lockhart SR, Pujol C, Soll DR. Opaque cells signal white cells to form biofilms in Candida albicans. EMBO J. 2006 May 17;25(10):2240-52.
- Huang G, Wang H, Chou S, Nie X, Chen J, Liu H. Bistable expression of WOR1, a master regulator of white-opaque switching in Candida albicans. Proc Natl Acad Sci U S A. 2006 Aug 22;103(34):12813-8.
- Huang G, Yi S, Sahni N, Daniels KJ, Srikantha T, Soll DR. N-acetylglucosamine induces white to opaque switching, a mating prerequisite in Candida albicans. PLoS Pathog. 2010 Mar 12;6(3):e1000806.
- Soll DR. Why does Candida albicans switch? FEMS Yeast Res. 2009 Oct;9(7):973-89.
- Srikantha T, Borneman AR, Daniels KJ, Pujol C, Wu W, Seringhaus MR, Gerstein M, Yi S, Snyder M, Soll DR. TOS9 regulates white-opaque switching in Candida albicans. Eukaryot Cell. 2006 Oct;5(10):1674-87.
- Zordan RE, Galgoczy DJ, Johnson AD. Epigenetic properties of white-opaque switching in Candida albicans are based on a self-sustaining transcriptional feedback loop. Proc Natl Acad Sci U S A. 2006 Aug 22;103(34):12807-12.
- Zordan RE, Miller MG, Galgoczy DJ, Tuch BB, Johnson AD. Interlocking transcriptional feedback loops control white-opaque switching in Candida albicans. PLoS Biol. 2007 Oct;5(10):e256.
- Soll DR. White-opaque switching in Candida albicans: cell biology, regulation, and function. Microbiol Mol Biol Rev. 2024 Jun 27;88(2):e0004322. doi: 10.1128/mmbr.00043-22.
- Du H, Ennis CL, Hernday AD, Nobile CJ, Huang G. N-Acetylglucosamine (GlcNAc) Sensing, Utilization, and Functions in Candida albicans. J Fungi (Basel). 2020 Aug 7;6(3):129.
- Qasim MN, Valle Arevalo A, Nobile CJ, Hernday AD. The roles of chromatin accessibility in regulating the Candida albicans white-opaque phenotypic switch. J Fungi (Basel). 2021 Jan 9;7(1):37.
PH RESPONSE
- Bensen ES, Martin SJ, Li M, Berman J, Davis DA. Transcriptional profiling in Candida albicans reveals new adaptive responses to extracellular pH and functions for Rim101p. Mol Microbiol. 2004 Dec;54(5):1335-51.
- Davis DA. How human pathogenic fungi sense and adapt to pH: the link to virulence. Curr Opin Microbiol. 2009 Aug;12(4):365-70.
- Davis D. Adaptation to environmental pH in Candida albicans and its relation to pathogenesis. Curr Genet. 2003 Oct;44(1):1-7. Erratum in: Curr Genet. 2003 Oct;44(1):58.
- Davis D, Wilson RB, Mitchell AP. RIM101-dependent and-independent pathways govern pH responses in Candida albicans. Mol Cell Biol. 2000 Feb;20(3):971-8.
- Kullas AL, Martin SJ, Davis D. Adaptation to environmental pH: integrating the Rim101 and calcineurin signal transduction pathways. Mol Microbiol. 2007 Nov;66(4):858-71.
- Ramon AM, Fonzi WA. Diverged binding specificity of Rim101p, the Candida albicans ortholog of PacC. Eukaryot Cell. 2003 Aug;2(4):718-28.
- Ramon AM, Porta A, Fonzi WA. Effect of environmental pH on morphological development of Candida albicans is mediated via the PacC-related transcription factor encoded by PRR2. J Bacteriol. 1999 Dec;181(24):7524-30.
QUORUM SENSING
- Kruppa M. Quorum sensing and Candida albicans. Mycoses. 2009 Jan;52(1):1-10.
- Langford ML, Atkin AL, Nickerson KW. Cellular interactions of farnesol, a quorum-sensing molecule produced by Candida albicans. Future Microbiol. 2009 Dec;4:1353-62.
- Roman E, Alonso-Monge R, Gong Q, Li D, Calderone R, Pla J. The Cek1 MAPK is a short-lived protein regulated by quorum sensing in the fungal pathogen Candida albicans. FEMS Yeast Res. 2009 Sep;9(6):942-55.
- Langford ML, Hasim S, Nickerson KW, Atkin AL. Activity and toxicity of farnesol towards Candida albicans are dependent on growth conditions. Antimicrob Agents Chemother. 2010 Feb;54(2):940-2.
STRESS RESPONSES
- Alonso-Monge R, Roman E, Arana DM, Pla J, Nombela C. Fungi sensing environmental stress. Clin Microbiol Infect. 2009 Jan;15 Suppl 1:17-9.
- Arana DM, Nombela C, Pla J. Fluconazole at subinhibitory concentrations induces the oxidative- and nitrosative-responsive genes TRR1, GRE2 and YHB1, and enhances the resistance of Candida albicans to phagocytes. J Antimicrob Chemother. 2010 Jan;65(1):54-62.
- Deveau A, Piispanen AE, Jackson AA, Hogan DA. Farnesol induces hydrogen peroxide resistance in Candida albicans yeast by inhibiting the Ras-cyclic AMP signaling pathway. Eukaryot Cell. 2010 Apr;9(4):569-77.
- Reedy JL, Filler SG, Heitman J. Elucidating the Candida albicans calcineurin signaling cascade controlling stress response and virulence. Fungal Genet Biol. 2010 Feb;47(2):107-16.
- Rodaki A, Bohovych IM, Enjalbert B, Young T, Odds FC, Gow NA, Brown AJ. Glucose promotes stress resistance in the fungal pathogen Candida albicans. Mol Biol Cell. 2009 Nov;20(22):4845-55.
- Berman J. Ploidy plasticity: a rapid and reversible strategy for adaptation to stress. FEMS Yeast Res. 2016 May;16(3):fow020.
THIGMOTROPISM, GALVANOTROPISM AND CONTACT-SENSING
- Sherwood J, Gow NA, Gooday GW, Gregory DW, Marshall D. Contact sensing in Candida albicans: a possible aid to epithelial penetration. J Med Vet Mycol. 1992;30(6):461-9.
- Gow NA, Perera TH, Sherwood-Higham J, Gooday GW, Gregory DW, Marshall D. Investigation of touch-sensitive responses by hyphae of the human pathogenic fungus Candida albicans. Scanning Microsc. 1994;8(3):705-10.
- Gow NA. Germ tube growth of Candida albicans. Curr Top Med Mycol. 1997 Dec;8(1-2):43-55.
- Brand A, Lee K, Veses V, Gow NA. Calcium homeostasis is required for contact-dependent helical and sinusoidal tip growth in Candida albicans hyphae. Mol Microbiol. 2009 Mar;71(5):1155-64.
- Brand A, Vacharaksa A, Bendel C, Norton J, Haynes P, Henry-Stanley M, Wells C, Ross K, Gow NA, Gale CA. An internal polarity landmark is important for externally induced hyphal behaviors in Candida albicans. Eukaryot Cell. 2008 Apr;7(4):712-20.
- Watts HJ, Very AA, Perera TH, Davies JM, Gow NA. Thigmotropism and stretch-activated channels in the pathogenic fungus Candida albicans. Microbiology. 1998 Mar;144 ( Pt 3):689-95.
Virulence
HOST-PATHOGEN INTERACTIONS
- Cockayne A, Odds FC. Interactions of Candida albicans yeast cells, germ tubes and hyphae with human polymorphonuclear leucocytes in vitro. J Gen Microbiol. 1984 Mar;130(3):465-71.
- Filler SG. Candida-host cell receptor-ligand interactions. Curr Opin Microbiol. 2006 Aug;9(4):333-9. Epub 2006 Jul 11.
- Gantner BN, Simmons RM, Underhill DM. Dectin-1 mediates macrophage recognition of Candida albicans yeast but not filaments. EMBO J. 2005 Mar 23;24(6):1277-86. Epub 2005 Feb 24.
- Gropp K, Schild L, Schindler S, Hube B, Zipfel PF, Skerka C. The yeast Candida albicans evades human complement attack by secretion of aspartic proteases. Mol Immunol. 2009 Dec;47(2-3):465-75. Epub 2009 Oct 31.
- Lorenz MC, Fink GR. Life and death in a macrophage: role of the glyoxylate cycle in virulence. Eukaryot Cell. 2002 Oct;1(5):657-62.
- McKenzie CG, Koser U, Lewis LE, Bain JM, Mora-Montes HM, Barker RN, Gow NA, Erwig LP. Contribution of Candida albicans cell wall components to recognition by and escape from murine macrophages. Infect Immun. 2010 Apr;78(4):1650-8. Epub 2010 Feb 1.
- Mochon AB, Ye J, Kayala MA, Wingard JR, Clancy CJ, Nguyen MH, Felgner P, Baldi P, Liu H. Serological profiling of a Candida albicans protein microarray reveals permanent host-pathogen interplay and stage-specific responses during candidemia. PLoS Pathog. 2010 Mar 26;6(3):e1000827.
- Mora-Montes HM, Bates S, Netea MG, Castillo L, Brand A, Buurman ET, Diaz-Jimenez DF, Jan Kullberg B, Brown AJ, Odds FC, Gow NA. A multifunctional mannosyltransferase family in Candida albicans determines cell wall mannan structure and host-fungus interactions. J Biol Chem. 2010 Apr 16;285(16):12087-95. Epub 2010 Feb 17.
- Rupp S. Interactions of the fungal pathogen Candida albicans with the host. Future Microbiol. 2007 Apr;2:141-51.
- Zhu W, Filler SG. Interactions of Candida albicans with epithelial cells. Cell Microbiol. 2010 Mar;12(3):273-82. Epub 2009 Nov 16.
- Brown AJ, Odds FC, Gow NA. Infection-related gene expression in Candida albicans. Curr Opin Microbiol. 2007 Aug;10(4):307-13.
- Gropp K, Schild L, Schindler S, Hube B, Zipfel PF, Skerka C. The yeast Candida albicans evades human complement attack by secretion of aspartic proteases. Mol Immunol. 2009 Dec;47(2-3):465-75.
- Hube B. Infection-associated genes of Candida albicans. Future Microbiol. 2006 Aug;1:209-18.
- Lorenz MC, Bender JA, Fink GR. Transcriptional response of Candida albicans upon internalization by macrophages. Eukaryot Cell. 2004 Oct;3(5):1076-87.
- Luo G, Ibrahim AS, Spellberg B, Nobile CJ, Mitchell AP, Fu Y. Candida albicans Hyr1p confers resistance to neutrophil killing and is a potential vaccine target. J Infect Dis. 2010 Jun 1;201(11):1718-28.
VIRULENCE AND VIRULENCE FACTORS
- Mitchell AP. Dimorphism and virulence in Candida albicans. Curr Opin Microbiol. 1998 Dec;1(6):687-92.
- Ibrahim AS, Magee BB, Sheppard DC, Yang M, Kauffman S, Becker J, Edwards JE Jr, Magee PT. Effects of ploidy and mating type on virulence of Candida albicans. Infect Immun. 2005 Nov;73(11):7366-74
- Wilson D, Thewes S, Zakikhany K, Fradin C, Albrecht A, Almeida R, Brunke S, Grosse K, Martin R, Mayer F, Leonhardt I, Schild L, Seider K, Skibbe M, Slesiona S, Waechtler B, Jacobsen I, Hube B. Identifying infection-associated genes of Candida albicans in the postgenomic era. FEMS Yeast Res. 2009 Aug;9(5):688-700.
- Wu W, Lockhart SR, Pujol C, Srikantha T, Soll DR. Heterozygosity of genes on the sex chromosome regulates Candida albicans virulence. Mol Microbiol. 2007 Jun;64(6):1587-604.
- Hogan DA, Sundstrom P. The Ras/cAMP/PKA signaling pathway and virulence in Candida albicans. Future Microbiol. 2009 Dec;4:1263-70.
- Kumamoto CA, Vinces MD. Contributions of hyphae and hypha-co-regulated genes to Candida albicans virulence. Cell Microbiol. 2005 Nov;7(11):1546-54.
- Lengeler KB, Tielker D, Ernst JF. Protein-O-mannosyltransferases in virulence and development. Cell Mol Life Sci. 2008 Feb;65(4):528-44.
- Mavor AL, Thewes S, Hube B. Systemic fungal infections caused by Candida species: epidemiology, infection process and virulence attributes. Curr Drug Targets. 2005 Dec;6(8):863-74.
- Monod M, Borg-von ZM. Secreted aspartic proteases as virulence factors of Candida species. Biol Chem. 2002 Jul-Aug;383(7-8):1087-93.
- Roman E, Arana DM, Nombela C, Alonso-Monge R, Pla J. MAP kinase pathways as regulators of fungal virulence. Trends Microbiol. 2007 Apr;15(4):181-90.
- Schaller M, Borelli C, Korting HC, Hube B. Hydrolytic enzymes as virulence factors of Candida albicans. Mycoses. 2005 Nov;48(6):365-77.
C. glabrata
C. parapsilosis
C. auris
C. dubliniensis
C. tropicalis
Current Taxonomy
The genus Candida was originally formed as a heterogeneous grouping of opportunistic pathogenic budding yeasts that lacked sexual reproduction (Groenewald et al., 2023). Further genomic study has revealed evolutionary distances between these species that argues for taxonomic revision within the budding yeast subphylum Saccharomycotina (Shen et al., 2020, Liu et al., 2024, Kidd et al., 2023). Interestingly, it appears pathogenicity within humans has evolved multiple times in this subphylum (Rokas, 2022)
NCBI has recently announced adoption of new taxonomy that affects the Candida community.
The specific name changes are as follows:
- Clavispora lusitaniae (syn. Candida lusitaniae )
- Pichia kudriavzevii (syn. Candida krusei )
- Meyerozyma guilliermondii (syn. Candida guilliermondii )
- Nakaseomyces glabratus (syn. Candida glabrata )
- Candidozyma auris (syn. Candida auris )
Strain Information
Candida albicans Strains
SC5314
Genotype: wild type
Notes: Wild-type strain used in the systematic sequencing project, the reference sequence stored in CGD. The original strain background from which most of the common laboratory strains are derived. This strain is virulent in a mouse model of systemic infection and is frequently used as a wild-type control. In their 2004 Genome Biology paper on C. albicans genome sequence, Frank Odds, Al Brown and Neil Gow explain the origins of SC5314: "Strain SC5314 was used in the 1980s by scientists at the E.R. Squibb company (now Bristol-Myers Squibb, see Note1) for their pioneering studies of C. albicans molecular biology. It was engineered by Fonzi and Irwin to provide the uridine autotrophic mutant that has been essential to most subsequent molecular genetic research into C. albicans. The strain is usually described merely as a 'clinical isolate', but it is worth setting on record that SC5314 was originally isolated from a patient with generalized Candida infection by Margarita Silva-Hutner (see Note 2) at the Department of Dermatology, Columbia College of Physicians and Surgeons (New York, USA). The original isolate number was 1775 and the strain is identical with strain NYOH#4657 in the New York State Department of Health collection. (This information was provided by Joan Fung-Tome at Bristol-Myers Squibb as a personal communication.) SC5314 belongs to the predominant clade of closely related C. albicans strains that represents almost 40% of all isolates worldwide, as determined by DNA fingerprinting and multi-locus sequence typing (A. Tavanti, A.D. Davidson, N.A.R.G., M.C.J. Maiden and F.C.O., unpublished observations)."
Note 1: The earliest publications that used SC5314 came in 1968 from Squibb Institute for Medical Research.
Note 2: The documentation of the work done by Margarita Silva-Hutner and her lab is preserved at Columbia University Archival Collections.
References: Odds et al., 2004. Genome Biol. 2004; 5(7): 230; Fonzi and Irwin, 1993. 1993 Jul;134(3):717-28; Aszalos et al., 1968. J Antibiot (Tokyo). 1968 Oct;21(10):611-5; Maestrone and Semar, 1968. Naturwissenschaften. 1968 Feb;55(2):87-8; Meyers et al., 1968. Appl Microbiol. 1968 Apr;16(4):603-8.
BWP17
Genotype: ura3::imm434/ura3::imm434 iro1/iro1::imm434 his1::hisG/his1::hisG arg4/arg4
Notes: Isogenic to the SC5314 strain. Uridine, histidine and arginine auxotroph derived from the RM1000 strain by deletion of the ARG4 gene. This strain has a heterozygous deletion on chromosome 5 that was inherited from the RM1000 parental strain.
References: Wilson et al., 1999. J Bacteriol. 1999 Mar;181(6):1868-74.
CAF2-1
Genotype: URA3/ura3::imm434 IRO1/iro1::imm434
Notes: URA3 heterozygous strain derived from the SC5314 strain. The 3-prime end of one copy of the IRO1 gene that resides adjacent to URA3 was inadvertently deleted during the construction of this strain. This strain is virulent in a mouse model of systemic infection and is frequently used as a wild-type control.
References: Fonzi and Irwin, 1993. Genetics 1993 Jul;134(3):717-28.
CAI4
Genotype: ura3::imm434/ura3::imm434 iro1/iro1::imm434
Notes: Isogenic to the SC5314 strain. Uridine auxotroph constructed by deletion of the second copy of URA3. The second copy of IRO1 was inadvertently deleted upon strain construction. As a result, the strain and its descendants have no functional copy of IRO1. This strain is avirulent in a mouse model of systemic infection unless complemented with URA3.
References: Fonzi and Irwin, 1993. Genetics 1993 Jul;134(3):717-28; Garcia et al., 2001. Yeast. 2001 Mar 15;18(4):301-11.
CAI8
Genotype: ura3::imm434/ura3::imm434 iro1/iro1::imm434 ade2::hisG/ade2::hisG
Notes: Isogenic to the SC5314 strain. Derived from the CAF2-1 strain by deletion of URA3 and both copies of ADE2 using the URA-blaster method.
References: Fonzi and Irwin, 1993. Genetics 1993 Jul;134(3):717-28.
P37005
Genotype: MTLa/MTLa
Notes: Wild-type clinical isolate. Naturally homozygous for the MTLa mating type locus.
References: Lockhart et al., 2002. Genetics. 2002 Oct;162(2):737-45.
Red3/6
Genotype: ade2/ade2
Notes: Isogenic to the WO-1 strain. Adenine auxotroph derived from the WO-1 strain by chemical mutagenesis using MNNG.
References: Srikantha et al., 1995. Mol Cell Biol. 1995 Mar;15(3):1797-805.
RM1000
Genotype: ura3::imm434/ura3::imm434 iro1/iro1::imm434 his1::hisG/his1::hisG
Notes: Isogenic to the SC5314 strain. Derived from the CAI4 strain by deletion of the HIS1 gene using the URA-blaster method (see Fonzi and Irwin, 1993 for details of this method). The standard RM1000 strain was found to have a heterozygous deletion on chromosome 5. RM1000#2 is an isolate that has been shown to have wild-type copies of chromosome 5.
References: Negredo et al., 1997. Microbiology. 1997 Feb;143 ( Pt 2):297-302.
SN87
Genotype: ura3::imm434::URA3/ura3::imm434 iro1::IRO1/iro1::imm434 his1::hisG/his1::hisG leu2/leu2
Notes: Isogenic to the SC5314 strain. Histidine and leucine auxotroph derived from the RM1000#2 strain by deletion of the LEU2 gene. This strain is virulent in a mouse model of systemic infection.
References: Noble and Johnson, 2005. Eukaryot Cell. 2005 Feb;4(2):298-309.
SN95
Genotype: ura3::imm434::URA3/ura3iro1IRO1/iro1his1his1arg4/arg4
Notes: Isogenic to the SC5314 strain. Histidine and arginine auxotroph derived from the RM1000#2 strain by deletion of the ARG4 gene. This strain is virulent in a mouse model of systemic infection.
References: Noble and Johnson, 2005. Eukaryot Cell. 2005 Feb;4(2):298-309.
SN152
Genotype: ura3/::imm434::URA3/ura3::imm434 iro1::IRO1/iro1::imm434 his1::hisG/his1::hisG leu2/leu2 arg4/arg4
Notes: Isogenic to the SC5314 strain. Histidine, leucine and arginine auxotroph derived from the RM1000#2 strain by deletion of the LEU2 and ARG4 genes. This strain is virulent in a mouse model of systemic infection.
References: Noble and Johnson, 2005. Eukaryot Cell. 2005 Feb;4(2):298-309.
SN250
Genotype: his1Δ/his1Δ, leu2Δ::C.dubliniensis HIS1 /leu2Δ::C.maltosa LEU2, arg4Δ /arg4Δ, URA3/ura3Δ::imm434, IRO1/iro1Δ::imm434
Notes: Isogenic to the SC5314 strain. Derived from SN87 by integration of C. dubliniensis HIS1 and C. maltosa LEU2 at the disrupted leu2 loci and then deleted for arg4. Derived from SN87 and QMY23.
References: Noble et al., 2010. Nat Genet. 2010 Jul;42(7):590-8;
Mitrovich et al., 2007. Genome Res. 2007 Apr;17(4):492-502.
WO-1
Genotype: MTLalpha
Notes: Wild-type clinical isolate that switches between white and opaque phenotypes at high frequency. The MTLa locus is absent in this strain (for more information see Lockhart et al., 2002). This strain has been sequenced by the Broad Institute (http://www.broadinstitute.org/annotation/genome/candida_group/GenomeDescriptions.html)
References: Slutsky et al., 1987. J Bacteriol. 1987 Jan;169(1):189-97; Lockhart et al., 2002. Genetics. 2002 Oct;162(2):737-45.
WUM5A
Genotype: MTLalpha/MTLalpha ura3-1Δ::FRT/ura3-2Δ::FRT
Notes: Isogenic to the WO-1 strain. Uridine auxotroph constructed by deletion of both copies of URA3.
References: Strauss et al. 2001. J Bacteriol. 2001 Jun;183(12):3761-9.
Candida glabrata (Nakaseomyces glabratus) Strains
B
Genotype: Wild type
Notes: Clinical isolate from a case of vaginitis that did not respond to fluconazole or boric acid treatment; it is virulent in a murine model of vaginitis. This strain was called LF 547.92 in the original publication.
References: Fidel et al., 1996. J Infect Dis. 1996 Feb;173(2):425-31.
BG2
Genotype: Wild type
Notes: Clinical isolate from a case of vaginitis that did not respond to fluconazole or boric acid treatment; it is virulent in a murine model of vaginitis
References: Cormack and Falkow, 1999. Genetics. 1999 Mar;151(3):979-87.
BG14
Genotype: ura3-delta(-285 +932)::Tn903NeoR
Notes: A ura3 derivative of the wild-type B2 clinical isolate.
References: Cormack and Falkow, 1999. Genetics. 1999 Mar;151(3):979-87.
BG462
Genotype: URA3
Notes: A derivative of the B2 clinical isolate with the URA3 gene restored
References: De Las Peñas, et al., 2003. Genes Dev. 2003 Sep 15;17(18):2245-58.
CBS138 (ATCC 2001)
Genotype: Wild type
Notes: Type strain for the Genolevures C. glabrata sequencing project. This strain has also been called NRRL-Y-65 and IFO 0622.
References: Dujon et al., 2004. Nature. 2004 Jul 1;430(6995):35-44; Koszul, et al., 2003. FEBS Lett. 2003 Jan 16;534(1-3):39-48.
2001T
Genotype: trp1
Notes: A trp1 auxotrophic strain derived from the CBS138 (ATCC 2001) wild-type strain.
References: Kitada et al., 1995. Gene. 1995 Nov 20;165(2):203-6.
2001HT
Genotype: his3 trp1::ScURA3
Notes: A his3 trp1 auxotrophic strain derived from 200T, a derivative of the CBS138 (ATCC 2001) wild-type strain.
References: Kitada et al., 1995. Gene. 1995 Nov 20;165(2):203-6.
NCCLS84 (ATCC 90030)
Genotype: Wild type
Notes: Wild-type strain
References: Espinel-Ingroff et al., 1992. J Clin Microbiol. 1992 Dec;30(12):3138-45
84u
Genotype: ura3
Notes: A ura3 derivative of the wild-type NCCLS84 (ATCC90030) strain.
References: Tsai et al., 2006. Antimicrob Agents Chemother. 2006 Apr;50(4):1384-92.
Candida parapsilosis Strains
ATCC 22019
Genotype: heterozygous MET/met
Notes: Application of parasexual genetic methods was used to determine that C. parapsilosis strain ATCC 22019 is is heterozygous MET/met
References: Whelan and Kwon-Chung, 1988. J Med Vet Mycol 1988 Jun;26(3):163-71.
CDC317
Genotype: Wild type
Notes: Reference strain used for the C. parapsilosis sequencing project. This isolate came from the hands of a hospital worker, who was the source for an outbreak of infection in a Mississippi community hospital in 2001.
References: Kuhn et al., 2004. Emerg Infect Dis. 2004 Jun;10(6):1074-81; Clark et al., 2004. J Clin Microbiol. 2004 Oct;42(10):4468-72.
CLIB214 (CBS 604; ATCC 22019; NRRL Y-12969)
Genotype: Wild type
Notes: Type strain, originally isolated from a patient with diarrhea in Puerto Rico by Ashford (1928). This strain is dependent on oxidative metabolism for growth since it lacks a fermentative pathway.
References: Ashford, 1928. Am J Trop Med 8:507�538. In: Kurtzman CP, Fell JW, Boekhout T (eds), 2011. The yeasts: a taxonomic study. p. 987-1278, 5th ed. Elsevier, London, United Kingdom; Logue et al., 2005. Eukaryot Cell 4(6):1009-17.
CPL2
Genotype: leu2Δ::FRT/leu2Δ::FRT, his1Δ::FRT/his1Δ::FRT, frt::CdHIS1
Notes: Leucine auxotroph laboratory strain
References: Németh et al., 2021. Virulence. 2021 Dec;12(1):937-950.
GA1
Genotype: Wild type
Notes: Wild-type clinical isolate. The SAT1 flipper method and the Candida albicans IMH3 gene have been used as dominant-selectable markers to construct gene knockouts in this strain.
References: Gácser et al., 2005. FEMS Microbiol Lett. 2005 Apr 1;245(1):117-21; Gácser et al., 2007. J Clin Invest. 2007 Oct;117(10):3049-58.
SR23 (CBS7157)
Genotype: ade- lys-
Notes: An adenine and lysine auxotroph isolated from a contaminated culture of Saccharomyces cerevisiae as a yeast with peculiar physiological features and carrying a linear mitochondrial DNA
References: Kovac et al., 1984. Mol Gen Genet 197:420-424; Nosek et al., 2004. Mol Genet Genomics 272(2):173-80; Mutalová et al., 2024. Microbiol Resour Announc. 2024 Jul 31;13(9):e00347-24.
Candida auris (Candidozyma auris) Strains
AR0382 (B11109)
Genotype: Wild type
Notes: Aggregative strain with high biofilm formation.
References: Vila et al., 2020. mSphere. 2020 Aug 5;5(4):e00760-20; Wang et al., 2024. Nat Commun. 2024 Oct 25;15(1):9212.
AR0387 (B8441)
Genotype: Wild type
Notes: Reference strain; non-aggregative strain with low biofilm formation.
References: Vila et al., 2020. mSphere. 2020 Aug 5;5(4):e00760-20; Wang et al., 2024. Nat Commun. 2024 Oct 25;15(1):9212.
AR0389
Genotype: erg11-Y132F
Notes: Has high drug resistance due to an ERG11 mutation (Y132F) and overexpression of the drug efflux transporter gene CDR1.
References: Shinohara et al., 2024. JAC Antimicrob Resist. 2024 Feb 21;6(1):dlad155.
AR0390 (B11205)
Genotype: erg11-K143R
Notes: Clinical isolate with high fluconazole resistance.
References: Rybak et al., 2021. Microbiol Spectr. 2021 Dec 8;9(3):e01585-21.
B11220
Genotype: Wild type
Notes: Clade II clinical isolate with susceptibility to all antifungal drugs.
References: Yang et al., 2024. Infect Immun. 2024 Jun 11;92(6):e0010324.
B11221
Genotype: Wild type
Notes: Clade III clinical isolate with resistance to caspofungin. Contains an L-rhamnose utilization cluster that appears to allow more ready use of alternate sugars, better drug tolerance, and better survival from host immune cell attack, possibly due to reduced β-(1,3)-glucans at the cell surface.
References: Yang et al., 2024. Infect Immun. 2024 Jun 11;92(6):e0010324.
Candida tropicalis Strains
ATCC 20336 (pK233)
Genotype: Wild type
Notes: This strain excretes alpha,omega-dicarboxylic acids as a by-product when cultured on n-alkanes or fatty acids as the carbon source.
References: Craft et al., 2003. Appl Environ Microbiol 69(10):5983-91.
ATCC 20913
Genotype: ura3/ura3
Notes: A uracil auxotroph derived from C. tropicalis ATCC 20336 by random mutagenesis.
References: Haas et al., 1990. J Bacteriol 172(8):4571-7.
ATCC 750
Genotype: Wild type
Notes: A wild-type fluconazole susceptible strain.
References: Barchiesi et al., 2000. Antimicrob Agents Chemother 44(6):1578-84.
NCYC2512
Genotype: Wild type
Notes: A wild-type saline soil isolate from Pakistan that is capable of producing large amounts of alpha,omega-dodecanedioic acid.
References: Rodriguez et al., 1996. Yeast 12(13):1321-9.
1230
Genotype: Wild type
Notes: A wild-type dicarboxylic acid-producing industrial strain.
References: He and Chen, 2005. Yeast 22(6):481-91.
Candida dubliniensis Strains
CD36
Genotype: Wild type
Notes: Type strain, oral isolate from an HIV+ patient in Ireland; also catalogued as NCPF 3949 and CBS-7987.
References: Sullivan et al., 1995. Microbiology (Reading) 1995 Jul:141 ( Pt 7):1507-21.
Protocols and Methods
Candida albicans Protocols and Methods
- Basic techniques
- Candida albicans: Methods and Protocols
- Cihlar and Calderone, 2009. Methods Mol Biol 2009:499:v
- Insertion mutagenesis
- Mielich et al., 2018 G3 (Bethesda) 2018 Mar 28;8(4):1139-1145
- CRISPR-Cas9 deletions
- Halder et al., 2019. Nat Protoc. 2019 Mar;14(3):955-975
- Shapiro et al., 2018. Nat Microbiol. 2018 Jan;3(1):73-82
- Wensing and Shapiro, 2022. Methods Mol Biol. 2022:2377:69-88
- Nguyen et al., 2021. Microbiol Spectr. 2021 Dec 22;9(3):e0182021
- 5-FOA assay for assessing mutation rates
- Lang and Murray, 2008. Genetics. 2008 Jan;178(1):67-82
- Virulence studies in Galleria mellonella
- Champion et al., 2016. Virulence. 2016 Oct 2;7(7):840-5
- Desbois and Coote, 2016. Virulence. 2016 Oct 2;7(7):840-5
- Junqueira, 2012. Virulence. 2012 Oct 1;3(6):474-6
- Wojda et al., 2020. Pathog Dis. 2020 Nov 23;78(9):ftaa057
Candida glabrata (Nakaseomyces glabratus) Protocols and Methods
- Basic techniques
- MIC assays
- Clinical and Laboratory Standards Performance Standards for Antifungal Susceptibility Testing of Yeasts, 3rd Edition
- Israel et al., 2021. J Clin Microbiol. 2021 Jun 18;59(7):e00311-21
- Genetic manipulation of C. glabrata
- Usher, 2024. Curr Protoc. 2024 Sep;4(9):e70014
- Transformation of C. glabrata
- Istel et al., 2015. Bio Protoc. 2015 Jul 20;5(14):e1528
- Istel et al., 2015. Bio Protoc. 2015 Jul 20;5(14):e1529
- CRISPR-Cas9-based deletions
- Enkler et al., 2016. Sci Rep. 2016 Oct 21:6:35766
- Maroc and Fairhead, 2019. Yeast. 2019 Dec;36(12):723-731
- Virulence studies in Galleria mellonella
- Champion et al., 2016. Virulence. 2016 Oct 2;7(7):840-5
- Desbois and Coote, 2016. Virulence. 2016 Oct 2;7(7):840-5
- Junqueira, 2012. Virulence. 2012 Oct 1;3(6):474-6
- Wojda et al., 2020. Pathog Dis. 2020 Nov 23;78(9):ftaa057
Candida parapsilosis Protocols and Methods
- Basic techniques
- Candida Species: Methods and Protocols
- Arévalo-Jaimes et al.,2023. Front Cell Infect Microbiol. 2023 Dec 13;13:1323619.
- Transformation of C. parapsilosis
- Nemeth et al.,2021. Virulence. 2021 Mar 17;12(1):937–950
- Zemanova et al.,2004. Curr Genet. 2004 Mar;45(3):183-6
- Virulence studies in Galleria mellonella
- Champion et al., 2016. Virulence. 2016 Oct 2;7(7):840-5
- Desbois and Coote, 2016. Virulence. 2016 Oct 2;7(7):840-5
- Junqueira, 2012. Virulence. 2012 Oct 1;3(6):474-6
- Wojda et al., 2020. Pathog Dis. 2020 Nov 23;78(9):ftaa057
- CRISPR-Cas9-based deletions
- Lombardi et al.,2022. mSphere. 2022 Dec 21;7(6):e0039322
- Lombardi and Butler,2022. Methods Mol Biol. 2022:2542:13-40
Candida auris (Candidozyma auris) Protocols and Methods
- Basic techniques
- Identification of C. auris
- Narayanan et al., 2022. Microbiol Spectr. 2022 Apr 27;10(2):e0063422
- Abdolrasouli and Fraser, 2022. Methods Mol Biol. 2022:2517:21-32
- Transformation of C. auris
- Ruiz and Lorenz, 2022. Methods Mol Biol. 2022:2517:95-110
- Kasir et al., 2022. Methods Mol Biol. 2022: 2517:89-94
- CRISPR-Cas9-based deletions
- Ennis et al., 2021. Microbiol Spectr. 2021 Dec 22;9(3):e0182021
- Nguyen et al., 2017. mSphere. 2017 Apr 26;2(2):e00149-17.
- Virulence studies in Galleria mellonella
- Borman et al., 2016. mSphere. 2016 Aug 18;1(4):e00189-16
- Garcia-Bustos et al., 2022. Emerg Microbes Infect. 2022 Dec;11(1):136-146
Candida dubliensis Protocols and Methods
- Basic techniques
- Identification
- Tintelnot at al., 2000. J Clin Microbiol. 2000 Apr;38(4):1599-608
- Bliss et al., 2003. J Clin Microbiol. 2003 Mar;41(3):1152-60
- Ells et al., 2011. Mycoses. 2011 Jan;54(1):1-16
- Kianipour et al., 2018. Adv Biomed Res. 2018 Apr 24:7:66
- Transformation and strain construction in C. dubliniensis
- Staib et al., 2000. Gene. 2000 Jan 25;242(1-2):393-8
- Staib et al., 2001. J Bacteriol. 2001 May;183(9):2859-65
- Virulence studies in Galleria mellonella
- Chupácová et al., 2018. Pathog Dis. 2018 Feb 1;76(1)
- Dekkerová-Chupáčová et al., 2018. Mycopathologia. 2018 Dec;183(6)
Candida tropicalis Protocols and Methods
- Basic techniques
- Transformation of C. tropicalis
- Rohrer and Picataggio, 1992. Appl Microbiol Biotechnol. 1992 Feb;36(5):650-4
- Xiang et al., 2014. Yi Chuan. 2014 Oct;36(10):1053-61
- CRISPR-Cas9-based deletions
- Li et al., 2022. Microbiol Spectr. 2022 Jun 29;10(3):e0005922
- Lombardi and Butler, 2022. Methods Mol Biol. 2022:2542:13-40
- Zhang et al., 2020. Biotechnol Bioeng. 2020 Feb;117(2):531-542
- Virulence studies in Galleria mellonella
- Mesa-Arango et al., 2013. Med Mycol. 2013 Jul;51(5):461-72
- Forastiero et al., 2013. Antimicrob Agents Chemother. 2013 Oct;57(10):4769-81
- García-Carnero et al., 2020. J Fungi (Basel). 2020 Aug 27;6(3):152
Species Comparisons by Topic
Ploidy
Notes: Candida albicans and its close relatives are all on a single phylogenetic branch characterized by diploidy, while the two typical haploid species (C. glabrata and C. auris) are on distinct, distantly related branches for which haploidy is the standard form. Note that the two haploid pathogens are more prone to pleiotropic resistance because only a single mutation is needed in the haploid genome to generate stable change.
References:
Typical diploids
- C. albicans
- C. dubliniensis
- C. parapsilosis
- C. tropicalis
Typical haploids
- C. glabrata
- C. auris
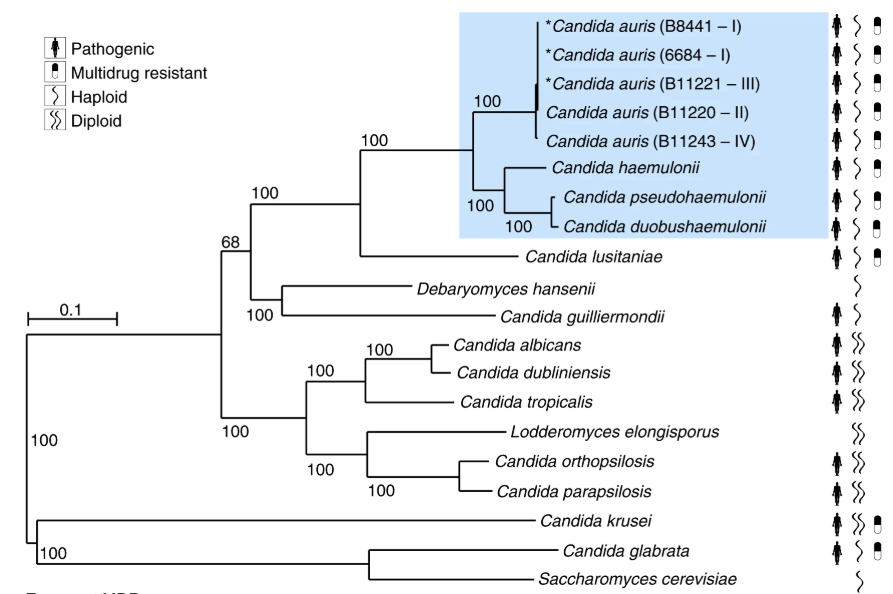
From Munoz et al., 2018
CUG codon usage
Notes: Yeasts of the Candida clade (as opposed to C. glabrata, also known as Nakaseomyces glabratus, which is on the Saccharomyces clade) have undergone an evolutionary reassignment of the CUG codon from leucine to serine. This is due to a novel serine-tRNA (ser-tRNACAG) that contains a guanosine at position 33, a position that is occupied by a pyrimidine (uridine) in nearly all other tRNAs.
References:
Reassigned CUG as serine
- C. albicans
- C. dubliniensis
- C. parapsilosis
- C. tropicalis
- C. auris
Standard CUG as leucine
- C. glabrata
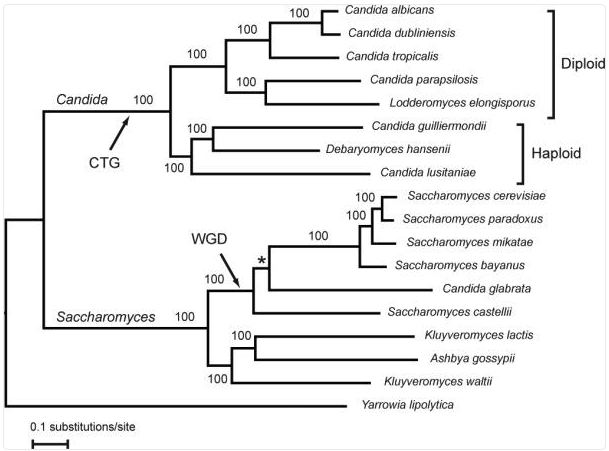
From Butler et al., 2009
Mitochondrial genome
Notes: Candida parapsilosis is unusual in having a mitochondrial genome that is linear and capped by telomeric ends. The close relatives of this species, C. orthopsilosis and C. metapsilosis and Candida subhashii, have a mix of circular and linear mitochondrial genomes. The other Candida spp. are exclusively circular.
References:
Exclusively linear
- C. parapsilosis
Mix of linear and circular
- C. orthopsilosis
- C. metapsilosis
- C. subhashii
Exclusively circular
- C. albicans
- C. dubliniensis
- C. tropicalis
- C. auris
- C. glabrata
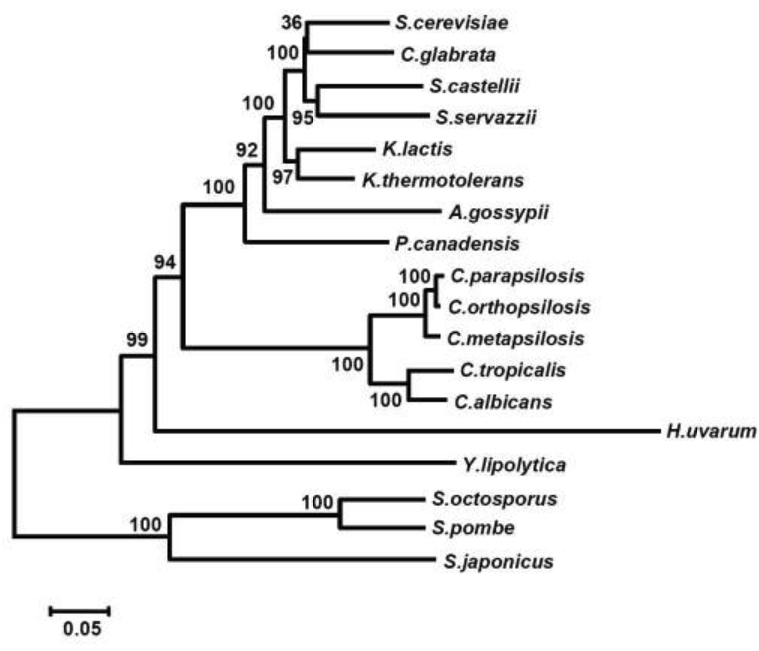
From Kosa et al., 2006
Filamentous growth
Notes: Candida spp. vary in filamentous growth as a means of infection/invasion. C. albicans and its close relative C. dubliniensis use true hyphae to penetrate tissues, while C. parapsilosis and C. tropicalis can penetrate tissues by means of pseudohyphae (lacking septa). In contrast, C. glabrata almost never forms filaments and infects in the yeast form. C. auris is an outlier in that it infects both by pseudohyphae and in the yeast form.
Note that the classifications below represent the typical forms for each species, but exceptions are not unusual.
References:
- Arita et al., 2022 (review)
- Kornitzer, 2019 (review)
- Silva et al., 2012 (review)
- Branco et al., 2023 (review)
- Bryak et al., 2024
- Groenewald et al., 2023
Invasive/infectious by true hyphae
- C. albicans
- C. dubliniensis
Invasive/infectious pseudohyphal growth (lacking septa)
- C. parapsilosis
- C. tropicalis
- C. auris
Invasive/infectious as yeast form
- C. glabrata
- C. auris
Biofilms
Notes: Candida species all form biofilms but not in the same manner. Biofilms all begin as adherence to a surface and then excretion of an extracellular matrix composed mainly of proteins, carbohydrates, and lipids. All species employ extracellular vesicles to facilitate excretion. They differ in the types of cells that inhabit the biofilm environment.
References:
- Zarnowski et al., 2022
- Kaur and Nobile, 2023
- Pujol et al., 2015
- Silva et al., 2009
- Singh et al., 2019
- Louvet et al., 2024
- Silva et al., 2012 (review)
Biofilms comprising primarily filamentous cells
- C. albicans
- C. dubliniensis (but mature biofilms are composed predominately of pseudohyphae rather than true hyphae (26432632)
Biofilms comprising mixed yeast and pseudohyphae
- C. tropicalis
- C. parapsilosis
- C. auris
Biofilms comprising primarily yeast cells
- C. glabrata
Pathogenicity
Notes: Candida species show heterogeneity in pathogenicity, where each species has a unique profile. See descriptions below.
References:
- Butler et al., 2009
- Casadevall et al., 2019
- Silva et al., 2012 (review)
- Branco et al., 2023 (review)
- Höfs et al., 2016 (review)
- Tóth et al., 2019 (review)
- Du et al., 2018
- Kołaczkowska and Kołaczkowski, 2016 (review)
C. albicans
- Exclusively commensal, C. albicans is fully adapted to the mammalian host, characterized by high thermotolerance and an advance ability to respond to changes in the body
- While non-albicans species are increasingly the cause of disease, C. albicans remains the most prevalent pathogen causing candidiasis
- Virulence properties are extensive, including the ability to (1) switch from the yeast form to the hyphal form; (2) secrete adhesins, biofilm matrix components, proteases, and hydrolytic enzymes; (4) undergo phenotypic change to allow sexual reproduction; (5) adapt metabolic processes to available nutrients; and (6) evade immune responses by various means
C. parapsilosis
- C. parapsilosis is the second or third most commonly isolated pathogenic species in several parts of the world
- Can live either commensally or freely in a wide range of ecological niches
- Has an enhanced ability to adhere to inert surfaces and is thus a particular problem in neonates, transplant recipients, and patients receiving parenteral nutrition
C. tropicalis
- Can live either commensally or freely in diverse environments
- More commonly associated with neutropenia and malignancy
C. glabrata
- Exclusively commensal, with an enormous ability to adapt to various stresses within the human body, including low availability of carbon, iron, and nitrogen, low pH, and both high and low oxygen levels
- Upon pathogenic invasion, can reach fatality levels of 50%
- More often associated with adults than children and is thus more prevalent in wealthier nations, where demographics skew to higher average age.
C. dubliniensis
- As a close relative of C. albicans, C. dubliniensis is a successful colonizer of the human host. However, it primarily lives as a benign commensal and only rarely causes invasive infection
C. auris
- C. auris was only recognized as a human pathogen in 2009 and may have only recently evolved the ability to colonize the human host
- Can live either commensally or freely in a wide range of ecological niches
- It has been hypothesized that C. auris made the shift to commensalism due to climate change, where increasing terrestrial temperatures made it more possible to bridge the transition into the high-temperature environment inside a mammal
- As C. auris preferentially colonizes the cooler skin rather than the hotter gut microbiome, new commensalism may be consistent with a recent acquisition of thermotolerance
- The inclination to colonize skin has made C. auris particularly problematic in hospital settings, where widespread outbreaks have become common
Mating/Reproduction
Notes: Candida species show heterogeneity in mating/reproduction, as described below. References are by species.
C. albicans
- Undergoes both heterothallic and homothallic reproduction
- Heterothallic reproduction between two haploids of opposite mating type occurs at low frequency and requires a phenotypic switch from the white to the opaque state, where the opaque state is mating competent
- Does not appear to undergo mating type switching within a strain
- Once mating occurs, the fungus does not undergo a meiotic program but instead reduces chromosome number by chromosome loss
- Parasexual (i.e., homothallic) reproduction via the mating of two diploids is likewise followed by chromosome loss leading to aneuploidy
- Has a remarkable tolerance for aneuploidy
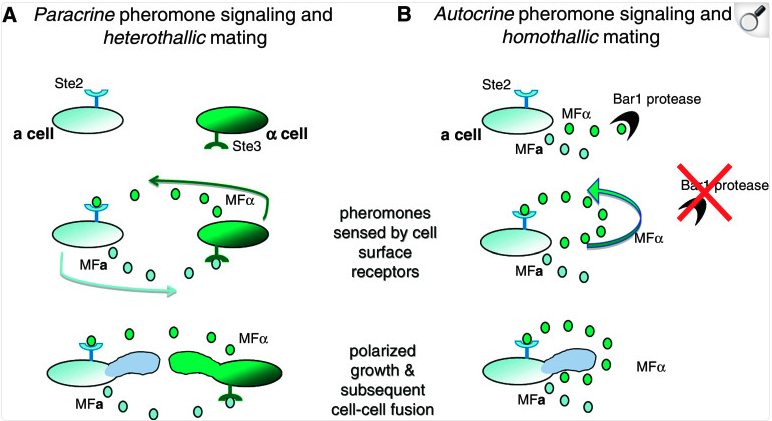
Slutsky et al., 1987 Alby and Bennett, 2010 (review) Butler et al., 2004 Noble and Johnson, 2007 (review)
C. dubliniensis
- Undergoes mating similarly to C. albicans and is likely to complete the mating cycle by parasexual loss of chromosomes rather than meiosis
- Requires phenotypic switching from white to opaque before mating
- Has a higher percentage (~30%) of natural strains that are MTL homozygous (a/a or alpha/alpha) than does C. albicans (~8%)
Pujol et al., 2004
Pujol et al., 2015
Alby and Bennett, 2010 (review)
Butler et al., 2004
C. tropicalis
- Undergoes both homothallic and heterothallic mating, followed by chromosome loss rather than meiosis
- Tetraploid progeny of homothallic matings can mate with diploid cells of the opposite mating type, generating unstable genomes that give rise to diverse offspring
Du et al., 2018
Alby and Bennett, 2010 (review)
C. parapsilosis
- Never observed to mate
Alby and Bennett, 2010 (review)
C. glabrata
- Never observed to mate
- However, there is genomic evidence for past recombination events and the genome contains the factors shown necessary for meiosis in the close relative S. cerevisiae
Dodgson et al., 2005
Bedekovic and Usher, 2023 (review)
C. auris
- Never observed to mate
- However, there is evidence in the genome for past recombination events, especially from before the clades split, and the species appears to contain the necessary factors for mating/meiosis
Drug resistance
Notes: Candida species show distinct drug resistance profiles. While all species can acquire resistance, inherent resistance and/or tolerance is especially concerning for C. auris and C. glabrata, likely due to their haploid genomes.
References:
- Carolus et al., 2021
- Kołaczkowska and Kołaczkowska, 2016
- Ahmady et al., 2024
- Pristov and Ghannoum, 2019
- Chowdhary et al., 2017
- Berman and Krysan, 2020
Concerning levels of inherent azole resistance
- C. glabrata
- C. auris
- C. krusei
Concerning levels of inherent echinocandin resistance
- C. glabrata
- C. auris
Concerning levels of amphotericin B resistance
- C. auris
Acquired resistance in nosocomial settings
- All species
Resistance within biofilms
- All species
Respiratory metabolism
Notes: The Crabtree effect explains why baker’s yeast is also called brewer’s yeast, in that S. cerevisiae preferentially converts sugar to alcohol in lieu of other types of respiration. This is referred to as Crabtree positive. Among the pathogenic yeasts, the only member with this ability is the phylogenetic outlier C. glabrata, which is more closely related to S. cerevisiae than to the other pathogenic yeasts. The other pathogenic yeasts are all Crabtree negative, meaning they do not produce alcohol as a product of respiration. For these non-fermentative yeasts, the ability to utilize many carbon sources (with little preference for glucose) is thought to provide an advantage for pathogenesis in various host niches.
References
Crabtree negative (non-fermentative)
- C. albicans
- C. dubliniensis
- C. parapsilosis
- C. tropicalis
- C. auris
Crabtree positive (fermentative)
- C. glabrata
Datasets Available in CGD
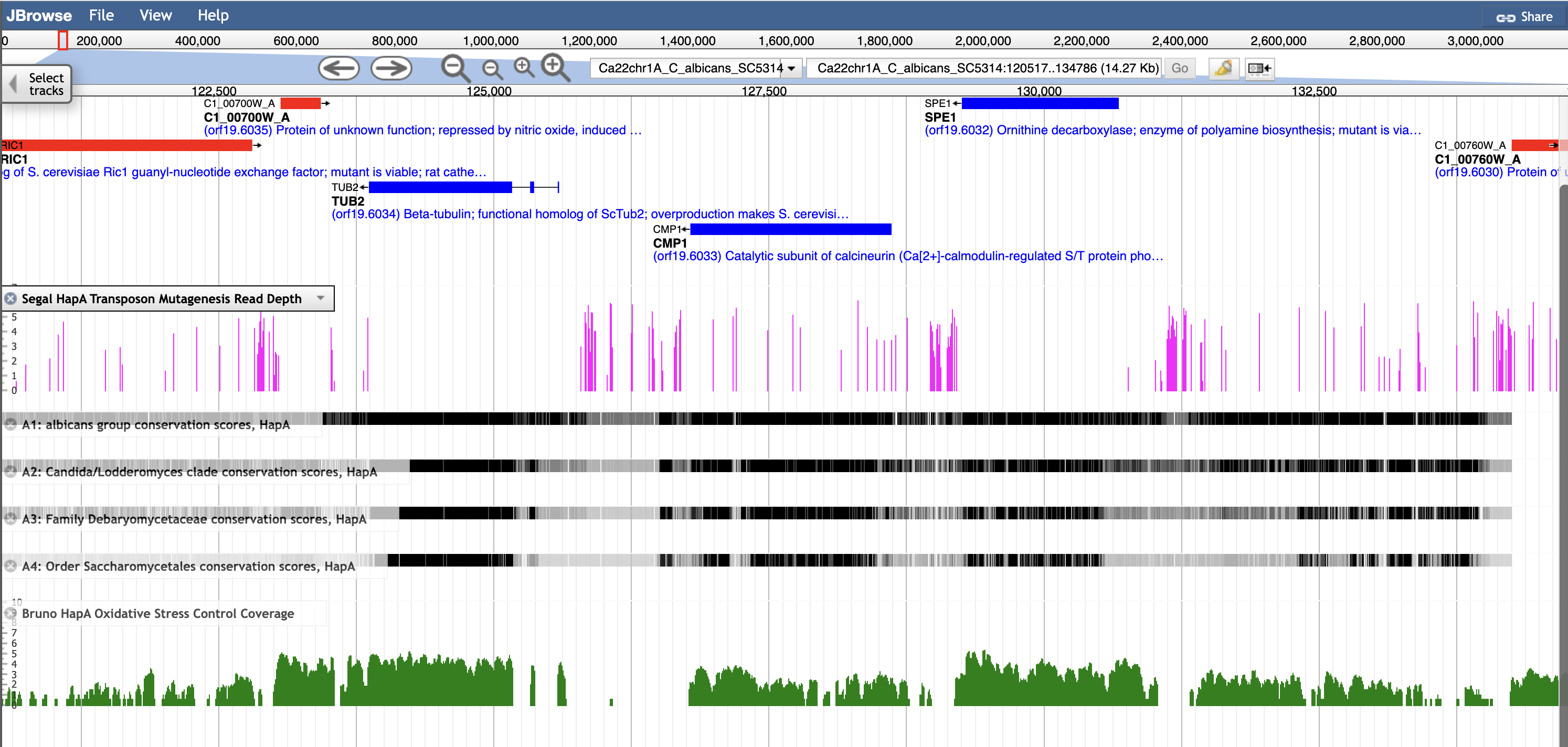
NOTE: To access a specific dataset in JBrowse, click "Select Tracks" at the upper left of the JBrowse page for the species and choose the dataset by first author name.
C. albicans in JBrowse
- Bruno et al., 2010. Comprehensive annotation of the transcriptome of the human fungal pathogen Candida albicans using RNA-seq. Genome Res. 2010 Oct;20(10):1451-8.
- Butler et al., 2009. Evolution of pathogenicity and sexual reproduction in eight Candida genomes. Nature. 2009 Jun 4;459(7247):657-62.
- Desai et al., 2013. Regulatory role of glycerol in Candida albicans biofilm formation. mBio. 2013 Apr 9;4(2):e00637-12.
- Lohse and Johnson, 2016. Identification and characterization of Wor4, a new transcriptional regulator of white-opaque switching. G3 (Bethesda). 2016 Jan 15;6(3):721-9
- Muzzy et al., 2013. Assembly of a phased diploid Candida albicans genome facilitates allele-specific measurements and provides a simple model for repeat and indel structure. Genome Biol. 2013;14(9):R97.
- Niemec et al., 2017. Dual transcriptome of the immediate neutrophil and Candida albicans interplay. BMC Genomics. 2017 Sep 6;18(1):696.
- Segal et al., 2018. Gene essentiality analyzed by in vivo transposon mutagenesis and machine learning in a stable haploid isolate of Candida albicans. mBio. 2018 Oct 30;9(5):e02048-18.
- Xie et al., 2013. White-opaque switching in natural MTLa/α isolates of Candida albicans: evolutionary implications for roles in host adaptation, pathogenesis, and sex. PLoS Biol. 2013;11(3):e1001525.
- Glazier et al., 2023. The Candida albicans reference strain SC5314 contains a rare, dominant allele of the transcription factor Rob1 that modulates filamentation, biofilm formation, and oral commensalism. mBio. 2023 Oct 31;14(5):e0152123.
- Shivarathri et al., 2019. The fungal histone acetyl transferase Gcn5 controls virulence of the human pathogen Candida albicans through multiple pathways. Sci Rep. 2019 Jul 1;9(1):9445.
- Rai, et al., 2024. Metabolic reprogramming during Candida albicans planktonic-biofilm transition is modulated by the transcription factors Zcf15 and Zcf26. PLoS Biol. 2024 Jun 21;22(6):e3002693.
- Zhang, et al. 2024. DNA damage checkpoints govern global gene transcription and exhibit species-specific regulation on HOF1 in Candida albicans. J Fungi (Basel). 2024 May 29;10(6):387.
C. parapsilosis in JBrowse
- Connolly, et al., 2013. The APSES transcription factor Efg1 is a global regulator that controls morphogenesis and biofilm formation in Candida parapsilosis. Mol Microbiol. 2013 Oct;90(1):36-53.
- Guida et al., 2011. Using RNA-seq to determine the transcriptional landscape and the hypoxic response of the pathogenic yeast Candida parapsilosis. BMC Genomics. 2011 Dec 22:12:628.
C. dubliniensis in JBrowse
- Grumaz et al., 2013. Species and condition specific adaptation of the transcriptional landscapes in Candida albicans and Candida dubliniensis. BMC Genomics. 2013 Apr 2:14:212.
- Singh-Babakh et al., 2021. Lineage-specific selection and the evolution of virulence in the Candida clade. Proc Natl Acad Sci U S A. 2021 Mar 23;118(12):e2016818118.
C. glabrata in JBrowse
- Vu, et al., 2021. The Candida glabrata Upc2A transcription factor is a global regulator of antifungal drug resistance pathways. PLoS Genet. 2021 Sep 30;17(9):e1009582.
- Ni, et al., 2023. The regulatory subunits of CK2 complex mediate DNA damage response and virulence in Candida glabrata. BMC Microbiol. 2023 Oct 28;23(1):317.
- Kumar, et al., 2024. SWI/SNF complex-mediated chromatin remodeling in Candida glabrata promotes immune evasion. iScience. 2024 Mar 27;27(4):109607.
- Bhakt et al., 2022. The SET-domain protein CgSet4 negatively regulates antifungal drug resistance via the ergosterol biosynthesis transcriptional regulator CgUpc2a. J Biol Chem. 2022 Oct;298(10):102485.
- Linde et al., 2015. Defining the transcriptomic landscape of Candida glabrata by RNA-Seq. Nucleic Acids Res. 2015 Feb 18;43(3):1392-406.
C. auris in JBrowse
- Simm, et al., 2022. Disruption of iron homeostasis and mitochondrial metabolism are promising targets to inhibit Candida auris. Microbiol Spectr 2022 Apr 27;10(2):e0010022.
- Jakab, et al., 2021. Transcriptional profiling of the Candida auris response to exogenous farnesol exposure mSphere. 2021 Sep-Oct; 6(5): e00710-21.
- Biermann and Hogan, 2022 Transcriptional response of Candida auris to the Mrr1 inducers methylglyoxal and benomyl. mSphere 2022 Jun 29;7(3):e0012422.
- Balla, et al., 2023. Total transcriptome analysis of Candida auris planktonic cells exposed to tyrosol. AMB Express 2023 Aug 2;13(1):81.
- Jenull et al., 2021. Transcriptome signatures predict phenotypic variations of Candida auris. Front Cell Infect Microbiol 2021 Apr 14:11:662563.
- Chow et al., 2023. The transcription factor Rpn4 activates its own transcription and induces efflux pump expression to confer fluconazole resistance in Candida auris. mBio 2023 Nov 28;14(6):e0268823
- Shivarathri, et al., 2022. Comparative transcriptomics reveal possible mechanisms of amphotericin B resistance in Candida auris. Antimicrob Agents Chemother. 2022 Jun 21;66(6):e0227621.
- Pelletier, et al., 2024. Candida auris undergoes adhesin-dependent and -independent cellular aggregation. PLoS Pathog. 2024 Mar 11;20(3):e1012076